AI and Machine Learning in Modern Plant Breeding
Explore how AI and ML are transforming agriculture through genomic selection, yield prediction, and climate-smart breeding strategies to ensure food security.
Artificial Intelligence and Machine Learning
in Modern Plant Breeding
[Your Name]
MSc Genetics & Plant Breeding
[Institution Name]
March 2026
MSc Seminar Presentation
Section 1: Foundation
Introduction
Global Challenges
Food security threatened by growing population (10 billion by 2050), climate change, and shrinking arable land
Need for Speed
Conventional breeding takes 10–15 years per variety; urgent need for faster, smarter crop improvement
Limitations of Conventional Breeding
Low throughput, labor-intensive, dependent on breeder expertise, limited predictive ability
"We need to produce 70% more food by 2050 with fewer resources"
Section 1: Foundation
Evolution
of Plant Breeding
Conventional Breeding
Pre-1900s–1960s
Selection, hybridization, phenotype-based
Molecular Breeding
1980s–2000s
Marker-assisted selection, QTL mapping
Genomic Breeding
2000s–2015
Whole-genome sequencing, SNP arrays, GWAS
Digital/AI Breeding
2015–Present
Big data, ML models, predictive analytics
Current Era
The shift from observing plants to
predicting performance using data
Section 1: Foundation
What is Artificial Intelligence (AI)?
AI
is the simulation of human intelligence by computer systems — enabling machines to learn, reason, and make decisions.
Machine Learning (ML)
Systems that learn from data to improve performance without being explicitly programmed.
Deep Learning (DL)
Multi-layered neural networks that mimic the human brain for complex pattern recognition.
Natural Language Processing (NLP)
AI that understands and generates human language — used in literature mining, chatbots.
SECTION 1: FOUNDATION
What is
Machine Learning (ML)?
ML enables computers to learn patterns from data and make decisions — without being explicitly programmed for each scenario.
SUPERVISED LEARNING
Learns from labeled data (input → known output).
Yield prediction, disease classification
UNSUPERVISED LEARNING
Finds hidden patterns in unlabeled data.
Clustering genotypes, trait grouping
REINFORCEMENT LEARNING
Learns by trial and reward.
Autonomous planting robots, irrigation optimization
Like a student (ML model) learning from past exam papers (data) to predict future exam answers (new decisions)
Section 1: Foundation
Why AI/ML in Plant Breeding?
Handles Big Data
Processes millions of SNPs, phenotypic records, and environmental data simultaneously
Improved Accuracy
ML models outperform traditional statistical methods in trait prediction
Speed
Reduces selection cycles from years to months
Cost Reduction
Fewer field trials needed through in silico prediction
Scalability
Works across environments, crops, and breeding programs
Predictive Power
Predicts performance before planting — saving resources
AI/ML bridges the gap between genomics and field performance
Section 2: Data in Modern Breeding
Types of Data Used in
Modern Breeding
Single Nucleotide Polymorphisms (SNPs), SSR markers, whole-genome sequences, haplotype maps.
DNA sequencing platforms (Illumina, Oxford Nanopore)
Yield, plant height, days to flowering, disease resistance, grain quality.
Field trials, greenhouse experiments, sensor data
Temperature, rainfall, soil pH, humidity, solar radiation.
Weather stations, IoT sensors, remote sensing satellites
The real power of AI/ML lies in integrating all three data types
Section 2
Big Data
Challenges
VOLUME
Millions of SNP markers, thousands of field plots, petabytes of sensor imagery
VARIETY
Heterogeneous data types — genomic, phenotypic, environmental, image-based
VELOCITY
Real-time streaming from sensors, drones, satellite feeds
Data Integration:
Combining data from different platforms, formats, and scales remains a key bottleneck
Solving these challenges requires AI-powered pipelines and cloud computing
Section 3: ML Workflow
ML Workflow in Plant Breeding
Each step will be explored in detail in the following slides
Step 1: Data Collection
ML Workflow — Step 1 of 7
Field Trials
Multi-Location Trials
Yield Records
Agromorphological Traits
Genomic Sequencing
Genotyping-by-Sequencing (GBS)
SNP Chips & Microarrays
Whole Genome Resequencing (WGRS)
Garbage In, Garbage Out
Step 2: Data Cleaning & Preprocessing
ML Workflow — Step 2 of 7
1. Handling Missing Values
Imputation (mean/median/KNN), deletion of poor-quality markers, BEAGLE/IMPUTE2 for genomic imputation
2. Normalization & Scaling
Min-max normalization, Z-score standardization — ensures equal weight across traits
3. Outlier Detection
Identify and remove erroneous field data points using statistical methods (IQR, boxplots)
4. Data Encoding
Converting categorical traits (resistant/susceptible) to numerical form for ML algorithms
Clean data = better model performance.
Preprocessing can account for 60–70% of ML project time.
ML Workflow — Step 3 of 7
Step 3: Feature Selection
Identifying the most informative variables (markers/traits) that contribute to the prediction outcome, while discarding noise.
FILTER METHODS
Statistical tests — correlation, chi-square, ANOVA. Fast but ignores model interaction.
WRAPPER METHODS
Recursive Feature Elimination (RFE) — tries subsets of features with the model.
EMBEDDED METHODS
LASSO, Ridge Regression, Random Forest importance scores — built into the model.
Selecting key SNP markers from thousands, choosing relevant environmental covariates.
Feature selection reduces overfitting, speeds up training, and improves interpretability.
ML Workflow — Step 4 of 7
Step 4: Model Selection
No single model fits all problems — model choice depends on data type and objective
REGRESSION MODELS
Linear/Ridge/LASSO — simple, interpretable. Used for: yield prediction, continuous trait modeling
RANDOM FOREST (RF)
Ensemble of decision trees. Robust, handles non-linearity. Used for: disease classification, marker selection
SUPPORT VECTOR MACHINE (SVM)
Finds optimal hyperplane to classify data. Used for: disease detection, trait classification
NEURAL NETWORKS / DEEP LEARNING
Multi-layered networks for complex patterns. CNN for image analysis, RNN for time-series
ML Workflow — Step 5 of 7
Step 5: Model Training
Data is split: 70–80% Training Set | 20–30% Test Set
Model iteratively learns patterns from training data
Parameters are updated through optimization (e.g., gradient descent)
Training Dataset
Labeled genomic + phenotypic data used to teach the model
Learning Patterns
Model finds associations between markers/traits and target variable
Hyperparameter Tuning
Adjusting model settings (learning rate, depth) for optimal performance
Overfitting vs Underfitting
Balance between learning enough and not memorizing noise
Training is iterative — the model improves with each pass through the data (epoch)
ML Workflow — Step 6 of 7
A model that performs well only on training data is useless — validation tells the true story.
Cross-Validation
Data is partitioned into k subsets. The model is recursively tested on each fold to provide a robust, unbiased accuracy estimate (e.g., 5-fold, 10-fold CV).
Accuracy Metrics
Performance is quantified using robust statistical metrics such as R², RMSE for continuous traits, and AUC-ROC curves for classification tasks.
Prediction Ability
Measured as the fundamental statistical correlation between predicted and observed values — a critically important metric in genomic selection.
Independent Test Set
A pristine, completely unseen dataset serves as the ultimate test validation to unequivocally confirm the final model's true field generalizability.
Section 4: Applications in Plant Breeding
Application 1:
Genomic Selection (GS)
Genomic Selection uses genome-wide marker data to predict Genomic Estimated Breeding Values (GEBVs) for all individuals — even those without phenotypic records.
Historical Standard
Introduced by Meuwissen et al. (2001) — now standard in animal and plant breeding.
Statistical & ML Models
Uses RRBLUP, GBLUP, Bayesian methods, Random Forest, Neural Networks.
Accuracy Dependencies
Accuracy depends strictly on: training population size, marker density, and trait heritability.
<strong style="color: #a8e063;">GS can reduce the breeding cycle by 30–50%</strong> by eliminating phenotyping bottlenecks.
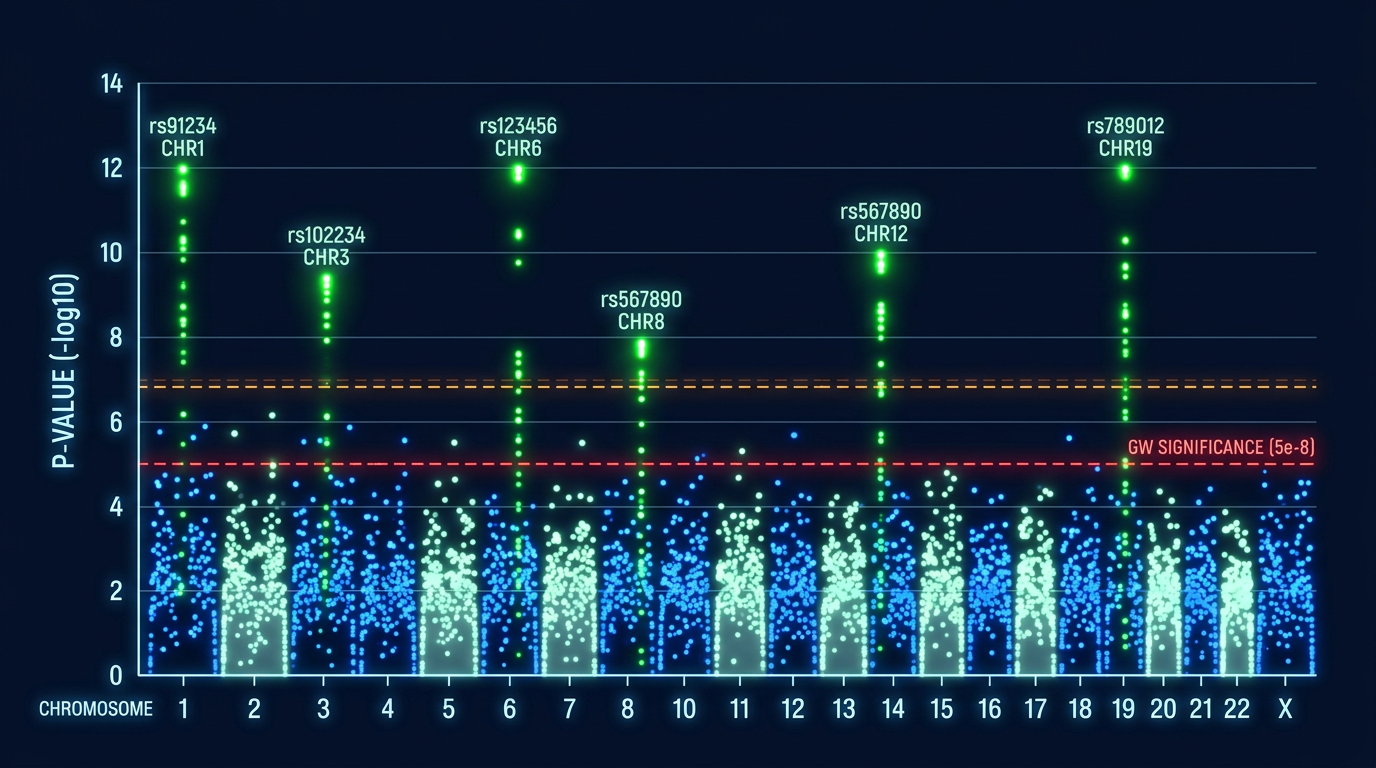
Genome-Wide Association Studies (GWAS) identify genomic regions (QTLs) significantly associated with traits of interest.
ML-GWAS in rice identified novel QTLs for drought tolerance and grain quality.
Random Forest GWAS, Deep Learning GWAS, Gradient Boosting
Higher Accuracy
ML handles population structure and epistasis better than conventional linear models.
Multi-trait Analysis
Simultaneously models multiple correlated traits.
Reduced False Positives
ML-based methods reduce spurious associations.
Non-linear Interactions
Captures epistatic effects missed by traditional ANOVA-based GWAS.
Application 3
Phenotyping
Automation
Image-Based Trait Analysis Using AI
Application 4
AI-Based
Disease Detection
Early, accurate, scalable detection using computer vision
Image Capture
Smartphone photos, drone imagery, field cameras capture leaf/plant images
Deep Learning (CNN)
Convolutional Neural Networks trained on thousands of disease images
Classification
Model identifies disease type, severity, spread stage
Alert & Recommendation
Farmer/breeder receives real-time alert with treatment advice
Wheat rust, rice blast, maize blight, powdery mildew
AI can detect disease 2–3 weeks before visible symptoms appear — saving entire crops
Application 5
Yield Prediction
Predicting crop performance before harvest using multi-environment data
Input Variables
Genotype (markers), environment (weather, soil), management practices, growth stage data
ML Models Used
Random Forest, Gradient Boosting (XGBoost), LSTM neural networks for time-series data
Model Output
Predicted yield per genotype per environment — enables best-fit variety recommendation
Performance
R² > 0.85
for yield prediction in maize and wheat
Real-World Application
IBM Watson Decision Platform for Agriculture predicts yield
3–4 weeks
before harvest with
90% accuracy
Yield prediction enables precision placement of varieties — right genotype, right environment
Application 6: Climate-Smart Breeding
Predicting Genotype × Environment Interaction (GEI) using AI
G×E Interaction
Same genotype performs differently in different environments — major challenge for breeders
ML Solution
Factored Regression, AMMI, Environmental Covariates with ML — models G×E accurately
Adaptability Prediction
Identify broadly adapted vs specifically adapted varieties
Climate Modeling
Project future climate scenarios → predict which genotypes will thrive in 2050
CIMMYT uses ML to identify drought and heat-tolerant wheat lines for future climate scenarios.
Sesame, maize, sorghum — C4 crops modeled under CMIP6 climate projections.
Climate-smart breeding = selecting today for tomorrow's climate
- artificial-intelligence
- machine-learning
- plant-breeding
- genomics
- agtech
- precision-agriculture
- crop-improvement
- data-science