Modeling Diabetic Vasculopathy with Blood Vessel Organoids
Learn how 3D human blood vessel organoids model diabetic microvasculopathy, identify NOTCH3 signaling drivers, and test new therapeutic interventions.
Human Blood Vessel Organoids as a Model of Diabetic Vasculopathy
An In-Depth Analysis of Wimmer et al. (Nature, 2019)
Journal Club Presentation
Agenda
<ul><li>Introduction: The Biomedical Problem</li><li>Central Research Question</li><li>Methods: Organoid Generation & Transplantation</li><li>Results: Modeling Diabetes & Mechanism</li><li>Therapeutic Intervention</li><li>Discussion: Strengths, Limitations & Alternatives</li></ul>
The Biomedical Problem: Diabetic Microvasculopathy
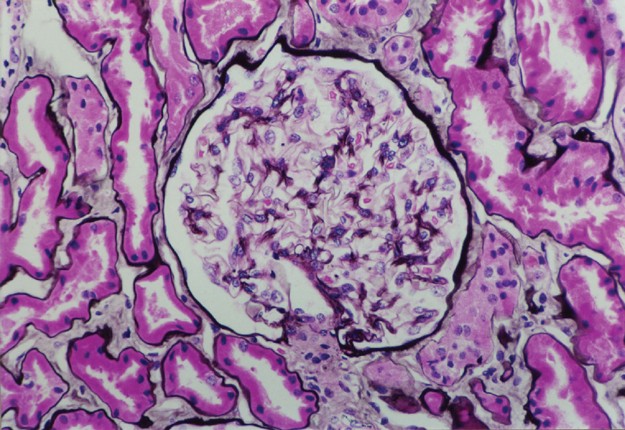
Diabetes is a major cause of blindness, kidney failure, and stroke. The hallmark pathology is changes in blood vessels, specifically the expansion of the basement membrane and loss of vascular cells. Current treatments often fail because we lack effective human models to study these early molecular changes.
Central Research Question
Can self-organizing 3D human blood vessel organoids, derived from pluripotent stem cells, faithfully model the structural and functional hallmarks of diabetic microvasculopathy?
Further, can this model identify key molecular drivers (like NOTCH3 signaling) and test potential pharmacological interventions?
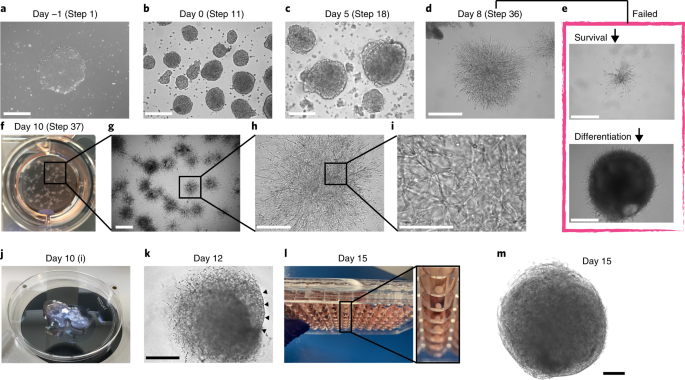
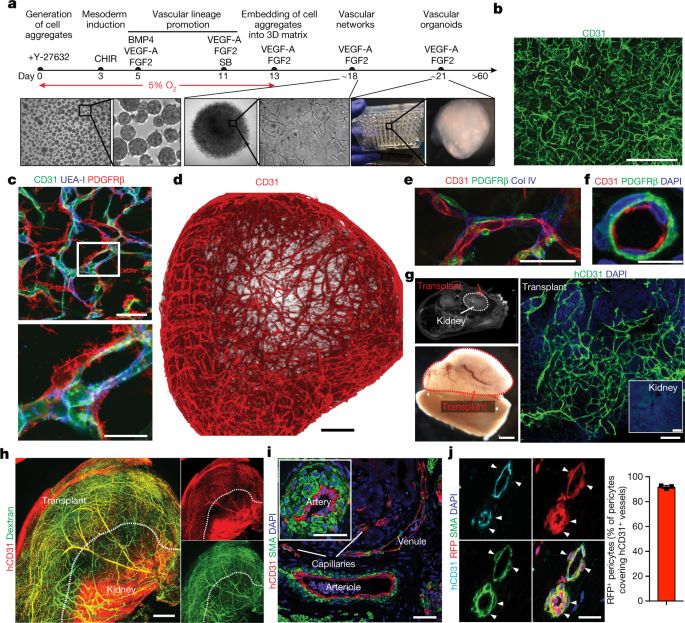
Methods: Generation of Human Blood Vessel Organoids
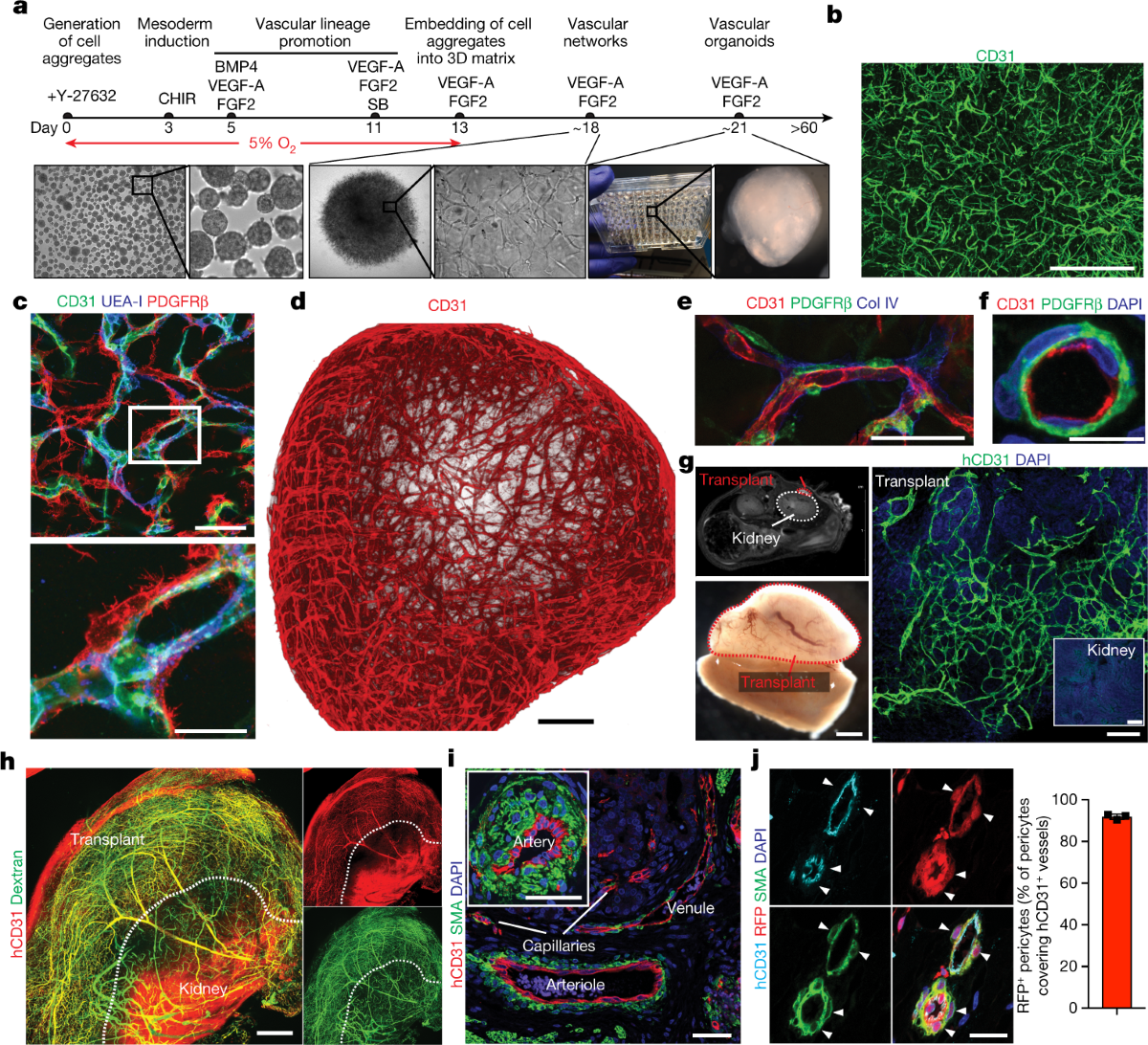
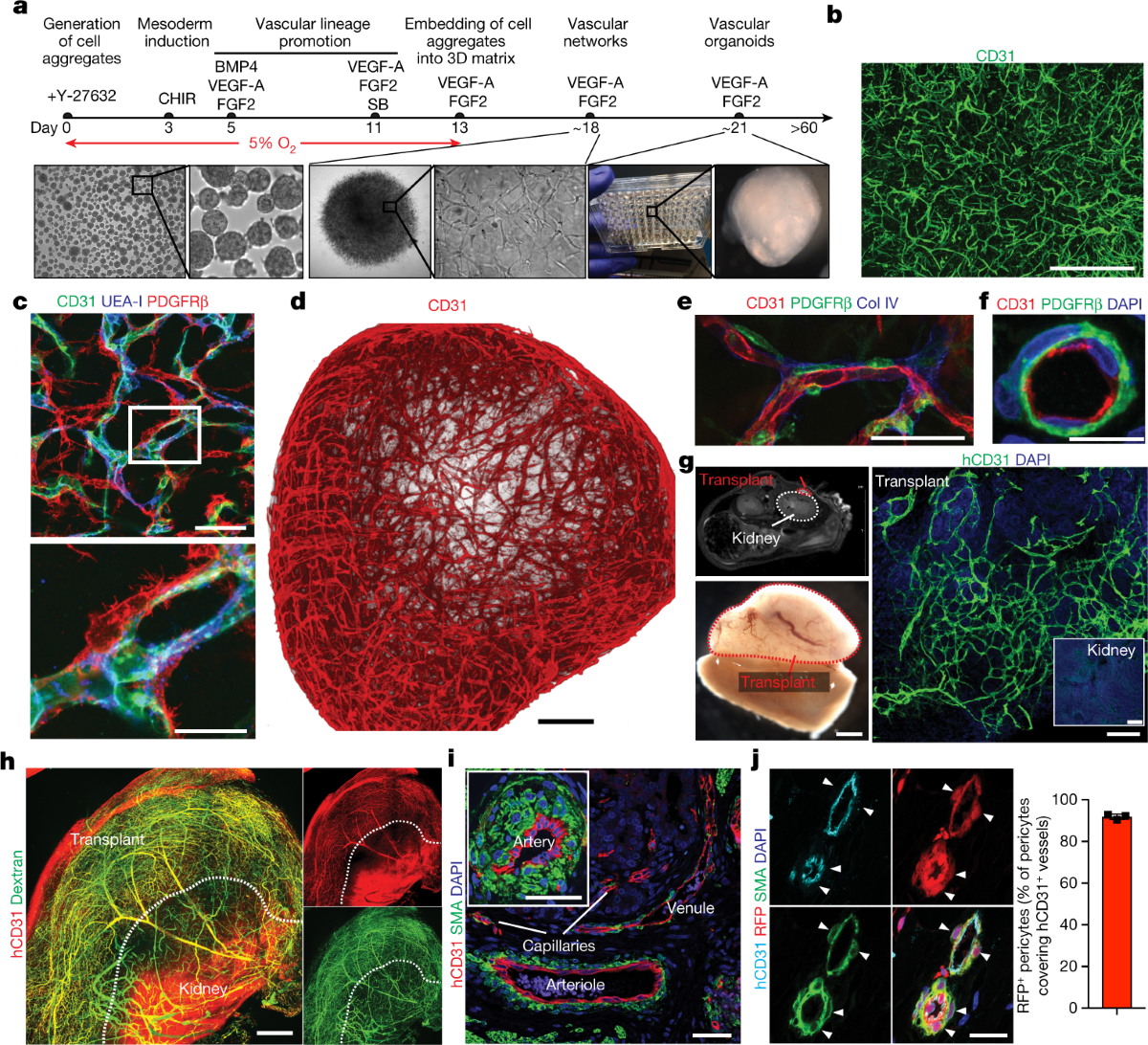
Protocol developed to differentiate iPSCs into vascular networks. Organoids self-assemble into capillaries enveloped by a basement membrane, containing both endothelial cells (CD31+) and pericytes (PDGFRβ+).
Study Design: In Vivo Transplantation
<ul><li><strong>Host:</strong> Immunodeficient NSG mice.</li><li><strong>Site:</strong> Transplanted under the kidney capsule.</li><li><strong>Result:</strong> Human vessels connect to mouse circulation (perfused).</li><li><strong>Diabetes Induction:</strong> Streptozotocin (STZ) treatment in mice to induce high blood sugar.</li></ul>
Results: Recapitulating Diabetic Microvasculopathy
<ul><li>In vitro organoids exposed to "Diabetic Milieu": High Glucose + TNF + IL-6.</li><li>Result observation: Massive thickening of the Basement Membrane (Collagen IV).</li><li>This mimics the exact pathology seen in human diabetic patients.</li></ul>
Quantifying Basement Membrane Thickening
Quantification of Collagen Type IV thickness significantly increases in diabetic conditions compared to controls.
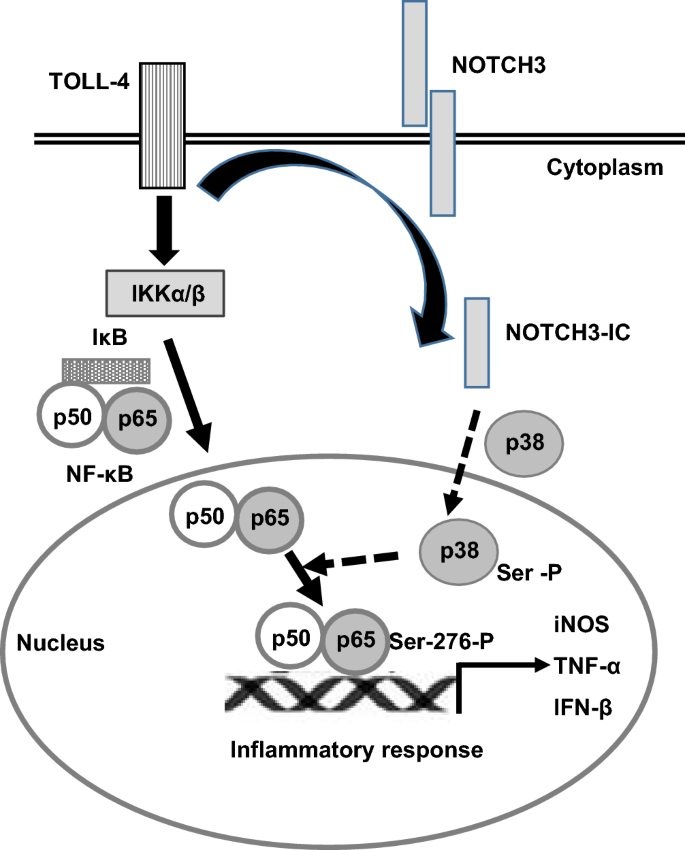
Mechanism: The Role of NOTCH3 and DLL4
<ul><li>The study identified <strong>NOTCH3</strong> and its ligand <strong>DLL4</strong> as key drivers of vasculopathy.</li><li>NOTCH3 is highly expressed in pericytes.</li><li>Blockade of NOTCH3 or DLL4 prevented basement membrane thickening.</li><li>Gene editing (CRISPR-Cas9) confirmed these findings.</li></ul>
Therapeutic Intervention: Gamma-Secretase Inhibition
DAPT (a γ-secretase inhibitor) blocks the cleavage required for Notch signaling. Treatment with DAPT successfully abrogated the expansion of Collagen IV and restored endothelial proliferation.
Critical Discussion: Strengths vs. Limitations
<strong>Strengths:</strong><br>• Human origin (overcomes species differences).<br>• 3D self-organization closely mimics in vivo architecture.<br>• Reproduces pathognomonic features (BM thickening) not seen in rodent models.
<strong>Limitations & Alternatives:</strong><br>• <em>In vitro</em> model lacks blood flow (shear stress).<br>• NSG mice lack a full immune system (inflammation may differ).<br>• <em>Alternative:</em> 'Organ-on-a-chip' could provide flow but lacks complex self-organization.
Conclusion & Takeaways
• Wimmer et al. established the first reliable human organoid model for diabetic microvasculopathy.<br>• They proved that Basement Membrane thickening is driven by the Diabetic Milieu via <strong>DLL4-NOTCH3 signaling</strong>.<br>• This model serves as a powerful platform for drug screening, identifying Gamma-Secretase inhibitors as a potential therapy.
- diabetic-vasculopathy
- organoids
- stem-cells
- microvasculopathy
- medical-research
- biotechnology
- notch3-signaling
- drug-screening