Basic Molecular Biology Techniques and DNA Analysis
Learn foundational molecular biology methods including DNA isolation, restriction enzymes, PCR, gel electrophoresis, and bioinformatics for genetic analysis.
Basic Molecular Biology Techniques
Methods for Analysis, Manipulation, and Separation of Nucleic Acids
Restriction Endonucleases
Type II restriction endonucleases are foundational tools in molecular biology. These enzymes recognize specific palindromic DNA sequences, typically 4–6 base pairs in length. Upon cleavage, they produce either 'blunt' ends or 'sticky' (staggered) ends. Staggered ends are particularly valuable because they can anneal to complementary strands, facilitating the construction of recombinant DNA.
DNA Isolation and Purification
Cell Lysis: Cells are ruptured gently (enzymatically or detergent-based) to prevent mechanical shearing of long DNA strands.
Contaminant Removal: RNA is degraded using RNase, while proteins are removed via phenol-chloroform extraction or proteinase K digestion.
Precipitation: Purified DNA is precipitated using absolute ethanol and then redissolved in TE buffer (Tris-EDTA) for storage.
Quality Check: Purity is verified using UV spectrophotometry or agarose gel electrophoresis.
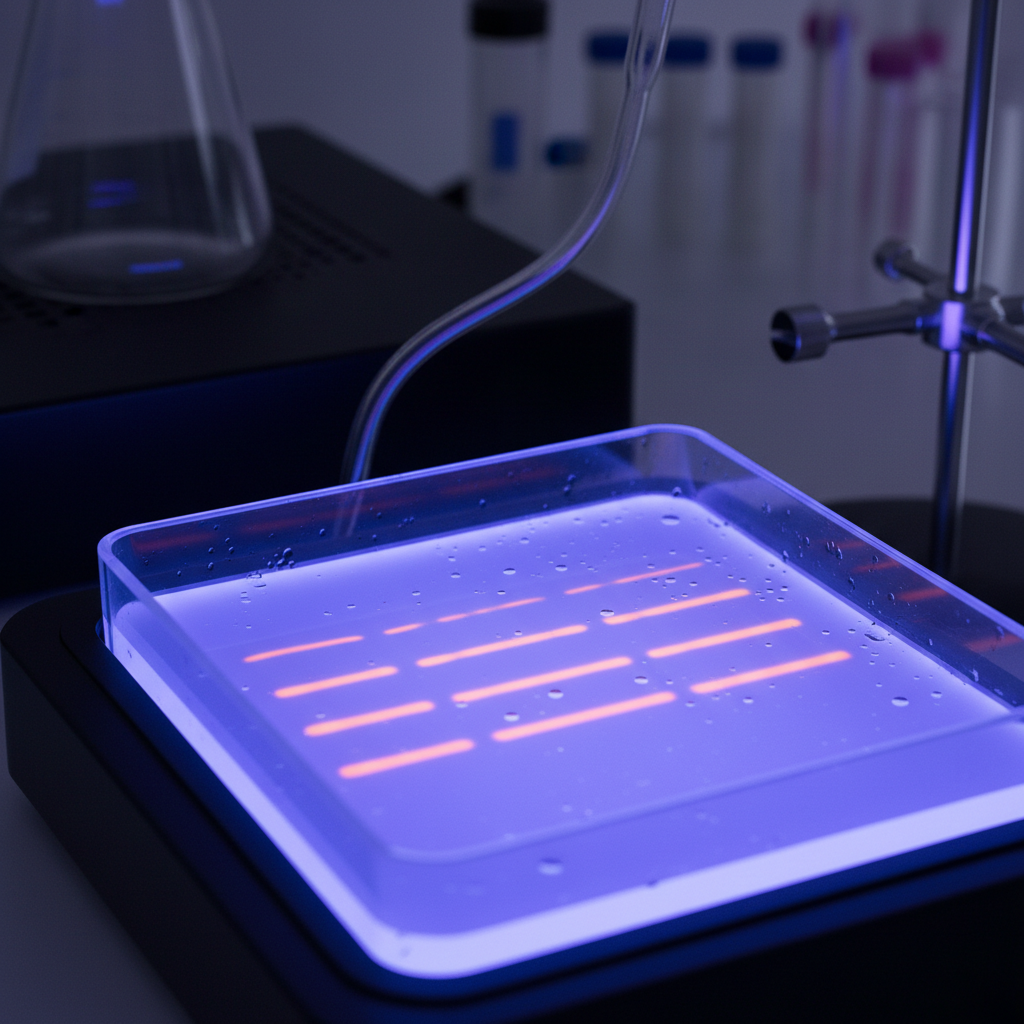
Gel Electrophoresis
Electrophoresis separates DNA molecules based on size within a matrix (agarose or polyacrylamide). Negatively charged DNA migrates toward the positive electrode, with smaller fragments moving faster. The DNA is typically stained with ethidium bromide, which intercalates between base pairs and fluoresces under UV light, allowing for visualization of the fragments perpendicular to the definition.
Restriction Mapping
Restriction mapping involves analyzing the size of fragments produced by digesting DNA with various restriction enzymes, both individually and in combination. By comparing fragment lengths (e.g., in a double digestion versus single digestions), the relative positions of restriction sites can be determined. This technique helps construct physical maps of DNA molecules and identify variations, such as Restriction Fragment Length Polymorphisms (RFLP).
Blotting Techniques
Southern Blotting: DNA is transferred from a gel to a nitrocellulose or nylon membrane via capillary action or electro-transfer, then hybridized with a labelled DNA probe.
Northern Blotting: Similar to Southern blotting but used for detecting and quantifying specific RNA transcripts (mRNA).
Ribonuclease Protection Assay: An alternative sensitive method for RNA analysis where unhybridized single-stranded RNA is digested, leaving the double-stranded probe-target complex.
Key Concept: 'Stringency' of hybridization (controlled by salt/temperature) determines the specificity of probe binding.
Spectrophotometric Constants
UV spectrophotometry is used to determine nucleic acid concentration. The concentration is calculated based on specific extinction coefficients at 260 nm (A260). A reading of 1 absorbance unit corresponds to different mass concentrations for DNA versus RNA molecules.
Polymerase Chain Reaction (PCR)
PCR allows for the rapid amplification of specific sequence fragments from a complex template. It requires knowledge of flanking sequences to design oligonucleotide 'primers'. By cycling temperatures (denaturation, annealing, extension), the target DNA increases exponentially. This technique eliminates the need for time-consuming cloning for many analytical applications.
Bioinformatics & Databases
Major DNA Databases: GenBank (USA), EMBL (Europe), DDBJ (Japan).
Protein Resources: Swiss-Prot and TREMBL provide curated protein sequence data (proteomics).
In Silico Research: The transition to computer-based analysis allows researchers to search for genes, predict structures, and analyze genomes without wet-lab experimentation initially.
Application: Crucial for designing gene probes and primers before physical synthesis.
Ideally, cell walls... should be digested enzymatically... and the cell membrane should be solubilised using detergent. Cell disruption should be kept to a minimum and should involve cutting or squashing of cells, rather than the use of shear forces.
Ralph Rapley - Basic Molecular Biology Techniques
Summary of Techniques
The modern molecular biology toolkit allows for the complete lifecycle of genetic analysis: from the careful isolation of nucleic acids and their digestion by restriction enzymes, to separation via electrophoresis and amplification by PCR. Coupled with bioinformatics, these methods enable the precise identification and manipulation of genetic material essential for biotechnology and medical research.
- molecular-biology
- dna-analysis
- pcr
- biotechnology
- genetics-research
- laboratory-techniques
- bioinformatics