Introduction to Graph Neural Networks for Drug Discovery
Learn how Graph Neural Networks (GNNs) process molecular data for drug discovery, including message passing, permutation invariance, and GCN architectures.
Graph Neural Networks: Learning Beyond the Grid
An intuition-first introduction into processing relational data, with a focus on Molecular Chemistry.
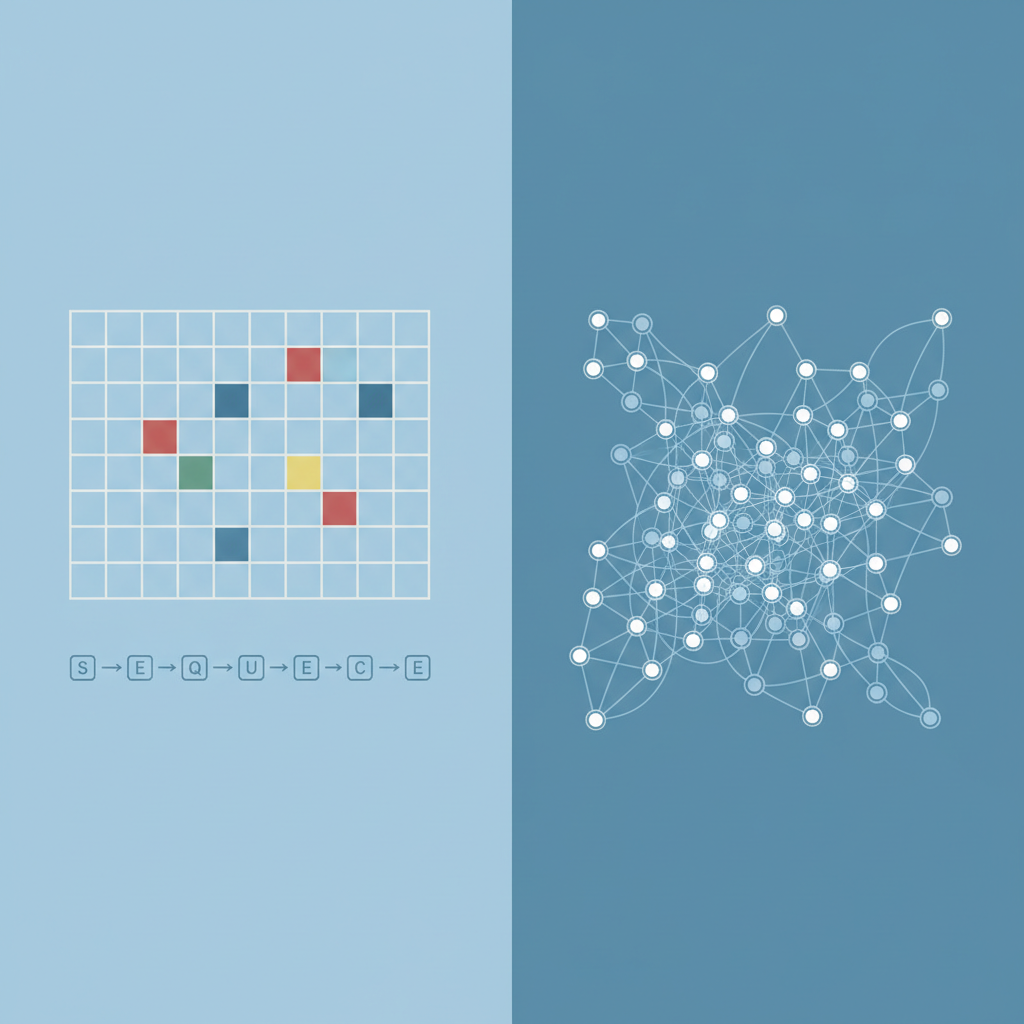
The Limitation of Traditional Deep Learning
Standard Neural Networks (CNNs, RNNs) expect data with a fixed structure, like grids (images) or sequences (text). They struggle when data has arbitrary size and complex topological connections.
The Real World is Relational: Molecular Example
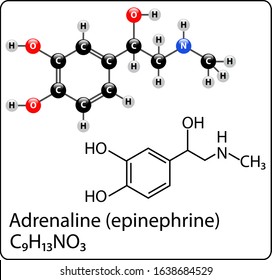
Running Case Study: Drug Discovery. Molecules are natural graphs.
Nodes = Atoms (Carbon, Oxygen, Nitrogen).
Edges = Chemical Bonds (Single, Double).
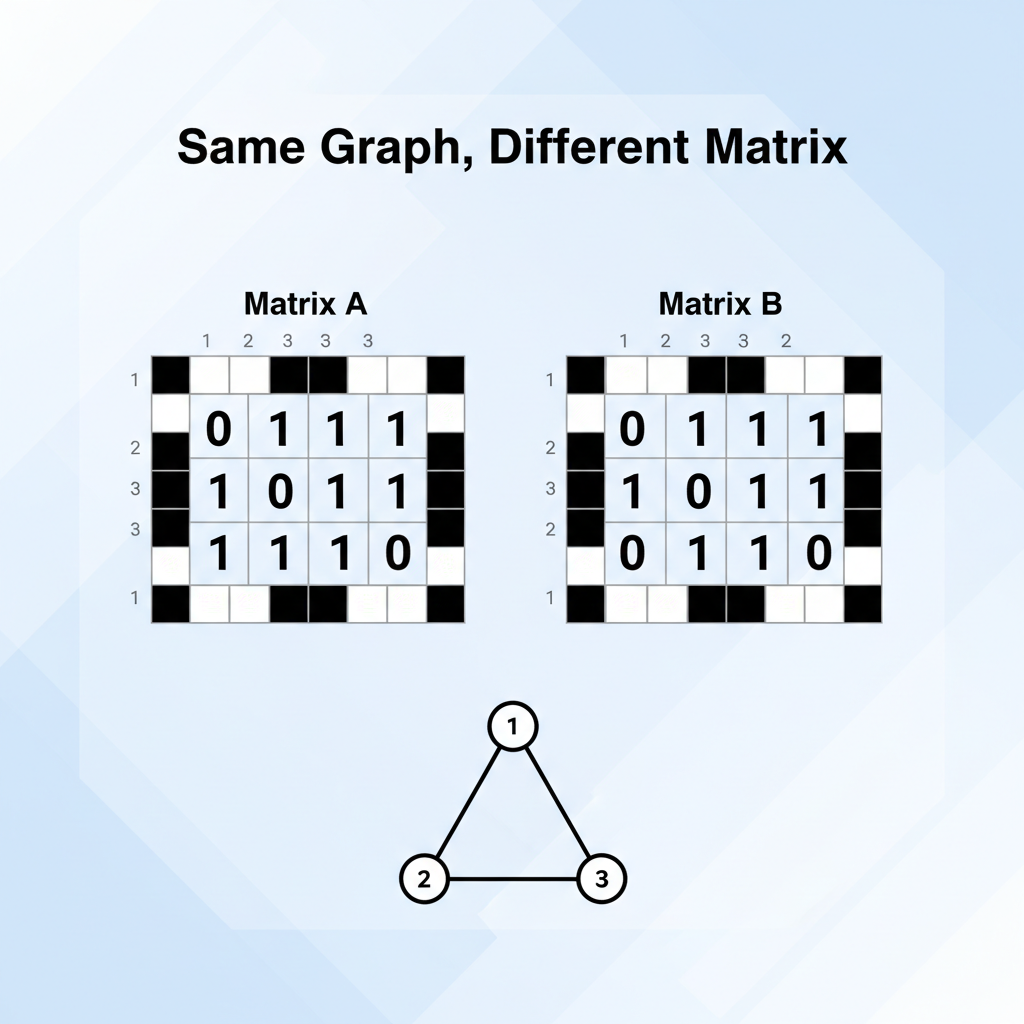
Problem: Permutation Invariance
Using a standard matrix (like an image) fails because graph nodes have no fixed order. If you label atoms differently (Atom 1 vs Atom 2), the matrix changes, but the molecule remains identical. A robust model must be 'permutation invariant'.
The GNN Solution: Neural Message Passing
Instead of processing the whole graph at once, we update each node based on its neighbors. This is local, scalable, and order-independent.
1. MESSAGE: Neighbors send their feature information.
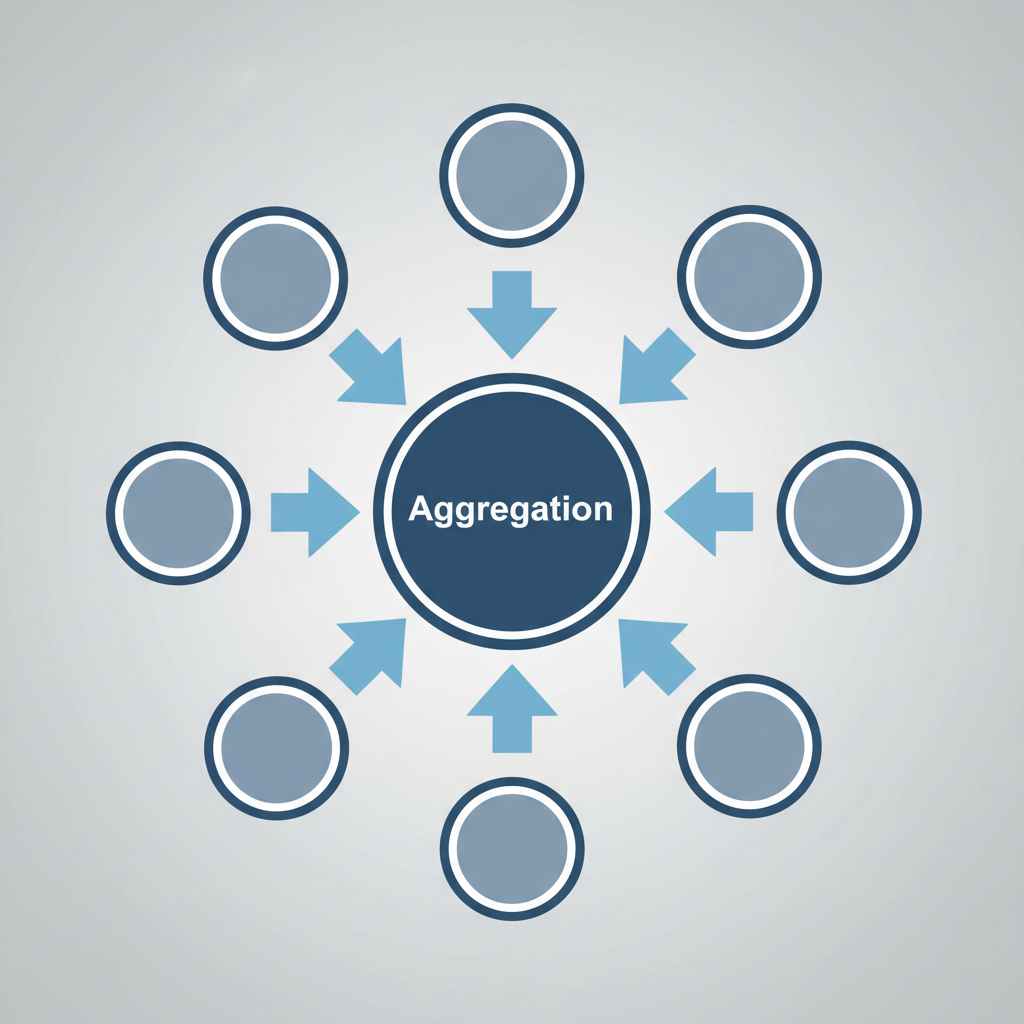
2. AGGREGATE: The node sums or averages these messages.
3. UPDATE: The node updates its own state using a Neural Network.
Under the Hood: The Aggregation Function
The state of node 'n' at time 't+1' depends on its previous state and the states of neighbors (ne[n]).
AGG: Sum, Mean, or Max pooling (permutation invariant).
W: Learnable weight matrix (shared across all nodes).
Scarselli et al. originally described this as an information diffusion mechanism to reach a stable state.
From Local Nodes to Global Prediction
Once message passing is done, every atom (node) has a vector representing its local chemical environment.
To classify the whole molecule (e.g., 'Is it Mutagenic?'), we use a READOUT function (sum or average of all node vectors) to get a single graph embedding.
Evolving Architectures
GCN (Graph Convolutional Network)
Uses a normalized weighted sum of neighbors. Simple and effective baseline.
GraphSAGE
Inductive learning. Samples a fixed number of neighbors (instead of all). Crucial for massive graphs (social networks) where nodes have thousands of connections.
GAT (Graph Attention Network)
Learns to weigh neighbors differently. A 'double bond' might be more important than a 'single bond'.
GNN Performance on Mutagenesis
Non-linear GNNs outperform other methods on graph-structured data sets (Mutagenesis). They successfully utilize the topological connections (bonds) that linear methods and standard feature-vector NNs struggle to capture fully.
Impact & Conclusion
GNNs have moved from theoretical curiosity to the backbone of modern AI in science.
Drug Discovery: predicting properties of molecules before synthesizing them.
Social Networks: Recommender systems (Pinterest/UberEats) use GraphSAGE.
Maps & Traffic: Predicting arrival times in Google Maps (ETA).
- gnn
- graph-neural-networks
- drug-discovery
- molecules
- machine-learning
- ai
- data-science