Catalytic Potential of Cupin-Type Enzymes (Tm1010, Tm1459)
Exploration of metal-dependent activity in cupin enzymes for biocatalysis, focusing on Mn²⁺ vs Cu²⁺ in HNL reactions and oxidative cleavage.
MASTER THESIS DEFENCE | BIOCATALYSIS
Catalytic Potential of Tm1010, Tm1459, and GtHNL in Biocatalytic Reactions
Effects of Copper on Expression and Various Reactions
Mathijs Zoon
5406889
Supervisors: U. Hanefeld, J. Berger | Faculty of Applied Sciences, TU Delft
April 2026
Faculty of Applied Sciences | Delft University of Technology
INTRODUCTION | BIOCATALYSIS
The World of Enzymes: Nature's Molecular Machines
Enzymes are biological catalysts that speed up reactions without being consumed
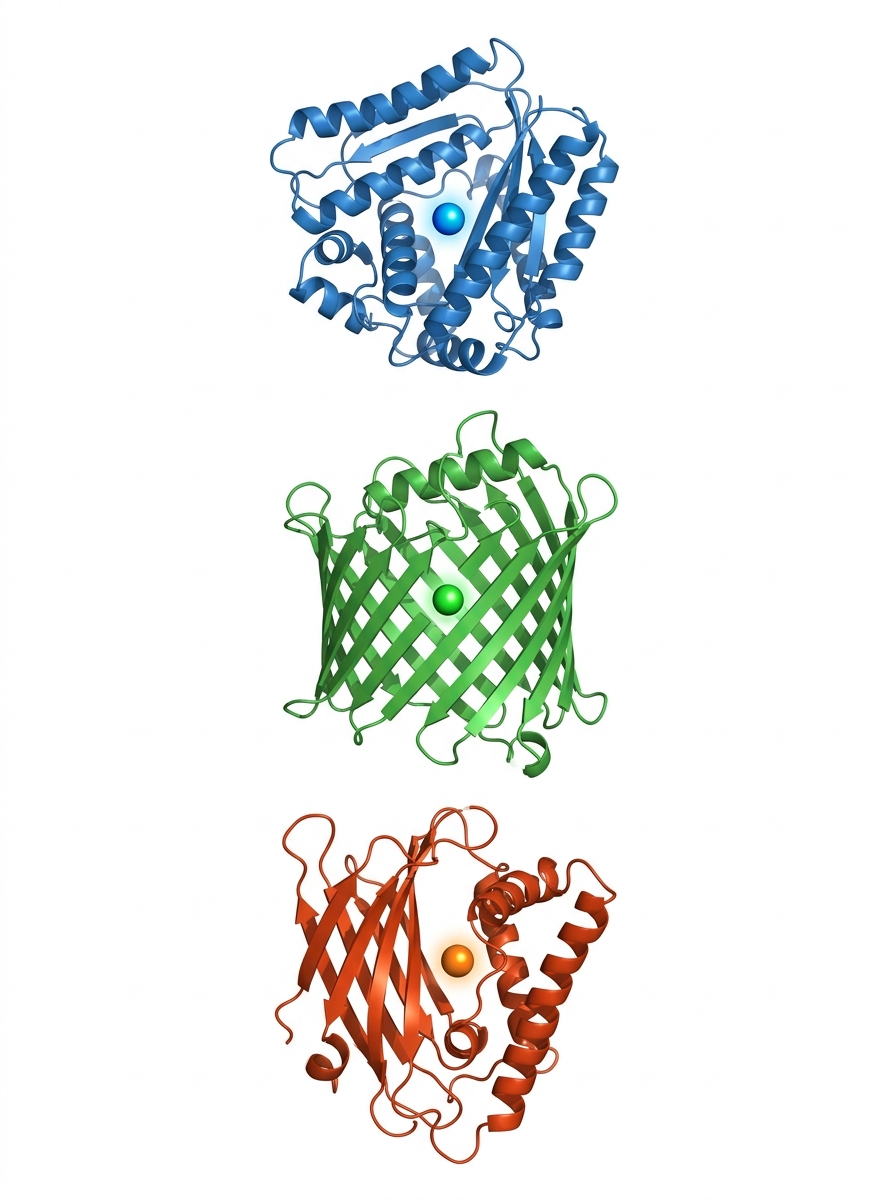
Cupin-type enzymes: a superfamily with a remarkable beta-barrel fold
Metal-dependent activity: manganese (Mn²⁺) and copper (Cu²⁺) as key cofactors
GtHNL-wt, Tm1010 & Tm1459: our three cupin heroes from this study
Why it matters
Enzymatic reactions occur under mild conditions — green chemistry for a sustainable future
Tm1010
Tm1459
GtHNL-wt
Faculty of Applied Sciences | Delft University of Technology
Reactions & Research Question: Cu²⁺ vs Mn²⁺
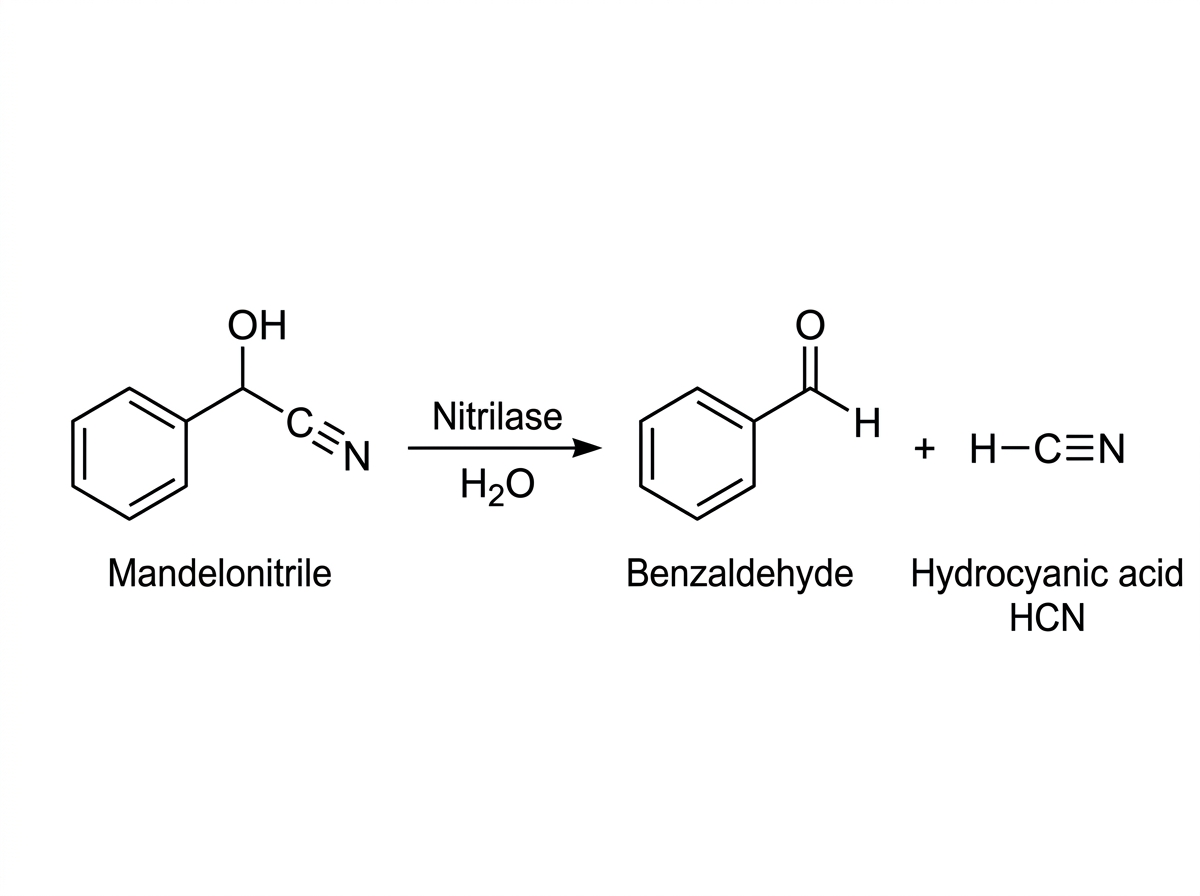
HNL Activity
Cleavage of cyanohydrins to aldehyde + HCN
Oxidative Cleavage
Alkene → carbonyl compounds (with TBHP)
Nitroaldol (Henry)
C-C bond formation, β-nitro alcohols
Michael Addition
Conjugate addition to α,β-unsaturated carbonyls
How does metal identity (Mn²⁺ vs Cu²⁺) and incorporation strategy affect the structural integrity and catalytic performance of Tm1010, Tm1459, and GtHNL-wt?
Motivated by: previous studies showing Tm1459 catalyzes oxidative cleavage (Hajnal et al. 2015) and copper-cupin systems enable non-native C-C bond forming reactions (Fujieda et al. 2020)
Methodology: How We Answer the Research Question
Each method was chosen to link enzyme structure to catalytic function
Structural Analysis
Metal Quantification: ICP-MS
Binding Thermodynamics: ITC
Catalytic Activity Assays
Faculty of Applied Sciences | Delft University of Technology
Two Strategies: Mn²⁺ vs Cu²⁺ Incorporation
Introduction — How We Make and Prepare the Enzymes
Both strategies were compared — Mn²⁺ expression + Cu²⁺ saturation was primarily used for reactions due to superior activity
Faculty of Applied Sciences | Delft University of Technology
RESULTS
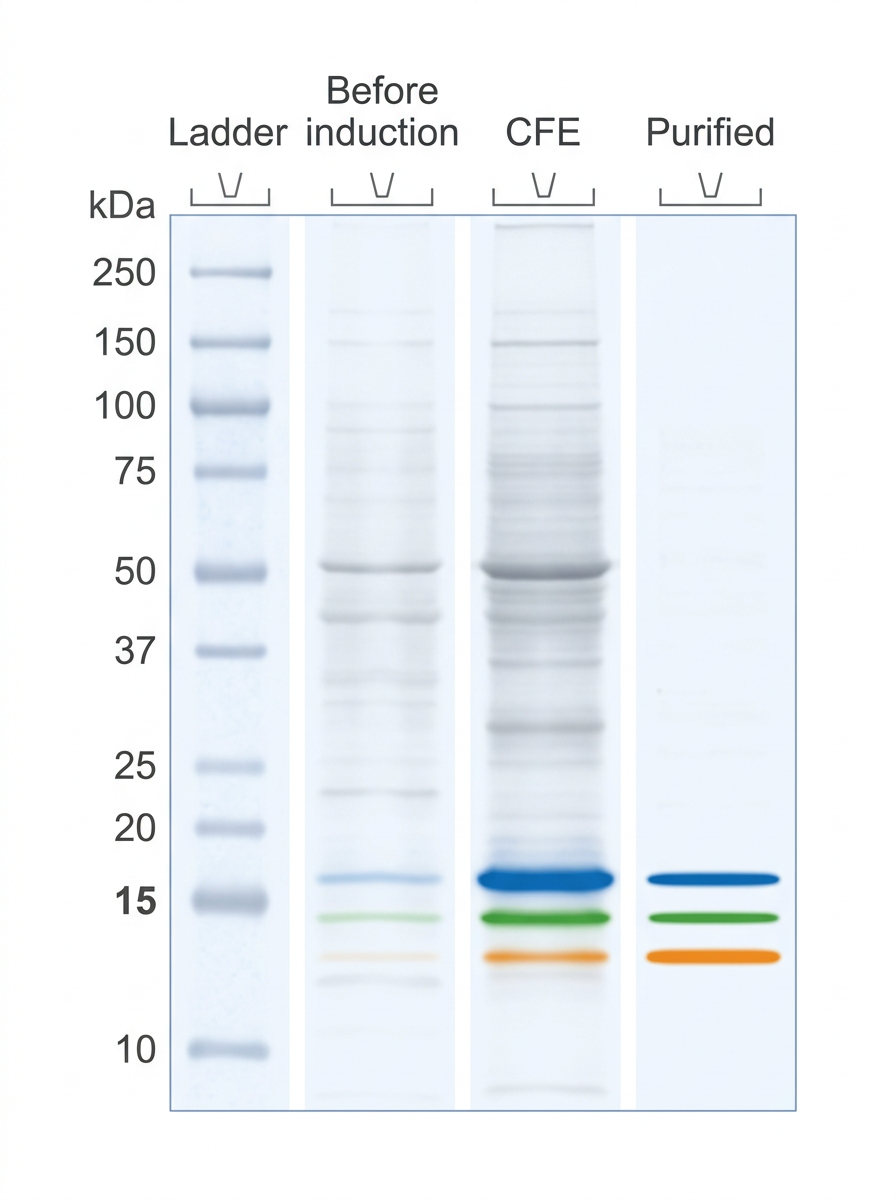
Results: Expression & Purification
All three enzymes (GtHNL-wt, Tm1010, Tm1459) successfully expressed in E. coli
Protein bands visible after IPTG induction; absent before induction
French Press (FP) method yielded purer samples than Heat Shock (HS) alone
FP: less protein loss in pellet fractions
Target protein bands: ~15-16 kDa for Tm1459, ~15 kDa for Tm1010 & GtHNL-wt
FP method selected for all subsequent copper expression experiments due to superior purity
Faculty of Applied Sciences | Delft University of Technology
RESULTS
Results: Dialysis & PD10 Desalting
Faculty of Applied Sciences | Delft University of Technology
RESULTS
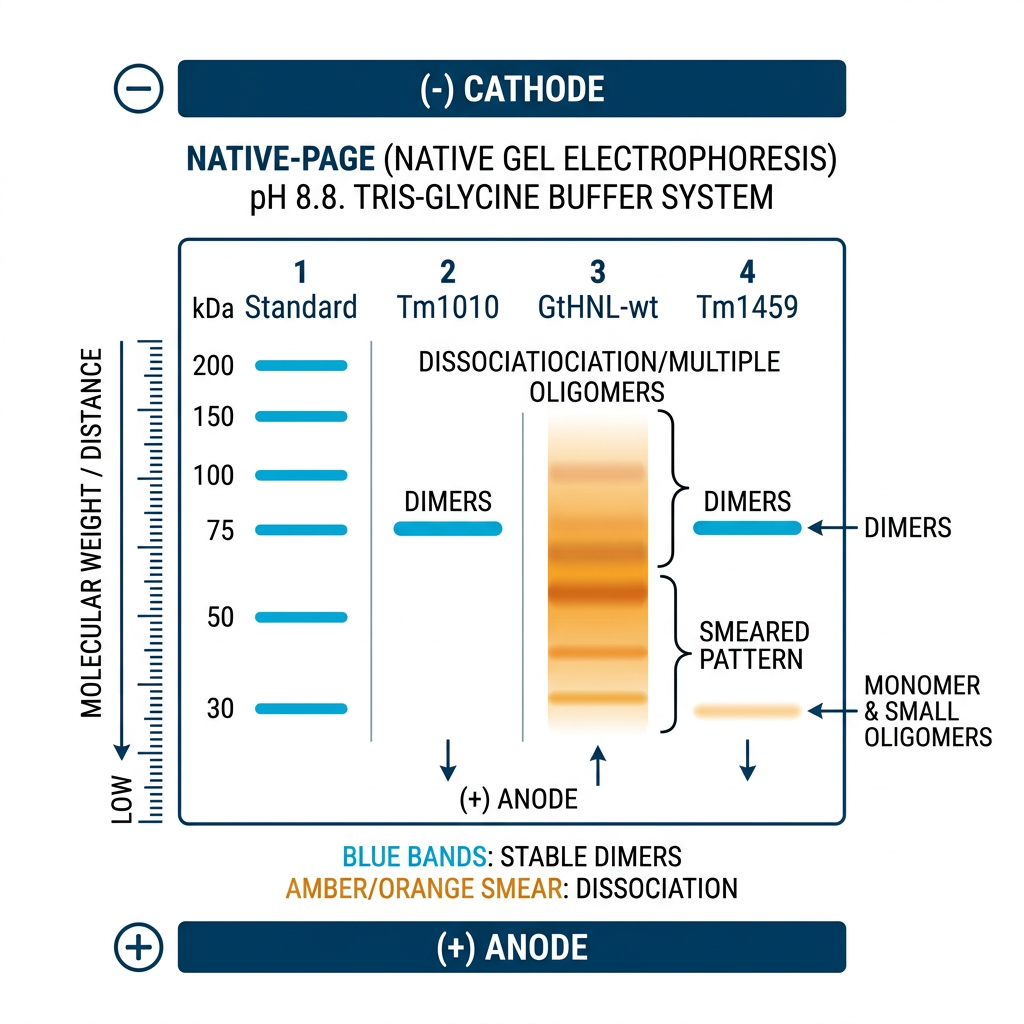
Results: Structural Analysis — Oligomeric State
SEC-HPLC Results
Tm1010
Apparent MW: 42.2 kDa | Monomer: 14.93 kDa
→ STABLE DIMER ✓
Tm1459
Apparent MW: 24.2 kDa | Monomer: 13.1 kDa
→ STABLE DIMER ✓
GtHNL-wt
Apparent MW: 19.0 kDa | Monomer: 14.29 kDa
→ PARTIAL DISSOCIATION ⚠️ (lower than expected for tetramer)
GtHNL-wt shows apparent instability under chromatographic conditions — dilution effects or protein-column interactions may cause dissociation. This may affect active site integrity.
Native PAGE Confirmation
Native PAGE confirms oligomeric assemblies — but GtHNL-wt discrepancy with SEC suggests dynamic dissociation in dilute conditions.
Key finding: Tm1010 & Tm1459 = stable dimers | GtHNL-wt = reduced oligomeric stability → impacts catalytic performance
Faculty of Applied Sciences | Delft University of Technology
RESULTS
Metal Binding — ICP-MS & ITC
Tm1459
Most defined metal binding. Enthalpy-driven Cu²⁺, entropy-driven Mn²⁺. Clear metalloenzyme behavior.
GtHNL-wt
Moderate, largely non-specific binding for both metals. Multiple surface sites.
Tm1010
Weak, poorly defined interactions for both metals. Lacks functional metal-binding site.
Structural stability (dimer) correlates with better-defined metal binding → better catalysis
Faculty of Applied Sciences | Delft University of Technology
Slide 9
RESULTS
Results: HNL Activity & Reverse HNL (Mandelonitrile Synthesis)
Faculty of Applied Sciences | Delft University of Technology
RESULTS
Results: Oxidative Cleavage (GC Analysis)
Faculty of Applied Sciences | Delft University of Technology
RESULTS
Non-Native Copper Reactions: Nitroaldol & Michael Addition
Nitroaldol (Henry) Reaction
Michael Addition
Faculty of Applied Sciences | Delft University of Technology
Slide 12
Conclusions & Recommendations
CONCLUSIONS
RECOMMENDATIONS
What Did We Find?
Where to Go Next?
Structural stability:
Tm1010 & Tm1459 = stable dimers. GtHNL-wt = partial dissociation → impacts catalytic performance.
Metal binding:
All enzymes bind Cu²⁺. Tm1459 shows most defined metal coordination. Tm1010 weakest binding.
Mn²⁺ preserves native HNL activity:
Cu²⁺ expression reduces HNL activity 4-10×. Mn²⁺-expression + Cu²⁺-saturation = optimal strategy.
Oxidative cleavage:
Tm1459-Mn²⁺ = 57% conversion (benchmark). Cu²⁺ generally reduces efficiency but timing of incorporation matters.
Nitroaldol (Henry):
Cu²⁺-bound enzymes achieve 60-70% conversion. Significant Cu²⁺ background reaction (~20%). Low enantioselectivity.
Michael addition:
Low conversion (<8%) across all systems. Protein scaffold enables moderate stereocontrol (up to 66% ee). Limited catalysis overall.
Overall:
Cu²⁺ unlocks non-native reactivity. Mn²⁺ = superior for native enzymatic function. Performance governed by structure, metal identity, and incorporation.
Optimize HNL conditions:
Use acidic pH (3.5-5) for reverse HNL to minimize background reaction and improve <i>ee</i>.
Improve immobilization:
Switch from Celite R-640 to R-633 (literature standard). Explore covalent immobilization to prevent leaching and deactivation.
Enhance enantioselectivity:
Protein engineering / directed evolution of active site residues (e.g., C106L in Tm1459). Focus on substrate positioning.
ITC optimization:
Improve Tm1010 binding experiments — signal quality was poor. Consider alternative buffer conditions.
Expand metal panel:
Test other divalent metals (Co²⁺, Zn²⁺, Ni²⁺) to further explore the catalytic versatility of cupin enzymes.
Flow chemistry:
Test Henry reaction in continuous flow with immobilized enzyme for improved process control.
Structural studies:
X-ray crystallography of Cu²⁺-loaded variants to directly visualize non-specific binding sites.
Faculty of Applied Sciences | Delft University of Technology
Thank You
Any Questions?
Mathijs Zoon
Master Thesis Biocatalysis | Student No: 5406889
Special thanks to U. Hanefeld, J. Berger, L. Koekkoek, M. Stampraat, S. Eustace, R. van Oosten, N. Karakitsou, and the entire BOC department
Faculty of Applied Sciences | Biocatalysis Department | Delft University of Technology | April 2026
- biocatalysis
- enzymes
- cupin-proteins
- green-chemistry
- biotechnology
- tu-delft
- metalloenzymes