Computational Pipelines for Protein Lipidation Prediction
Discover how bioinformatics and machine learning predict protein lipidation types like S-palmitoylation and myristoylation to map network biology.
A Computational Approach for Protein Lipidation
From Sequence Predictions to Network Biology
Bioinformatics Group Meeting
Scope of Study: Key Lipidation Types
S-Palmitoylation
Reversible attachment of palmitate to Cysteine residues via thioester linkage. Critical for dynamic membrane trafficking and signaling modulation.
N-Myristoylation
Irreversible co-translational attachment of myristate to an N-terminal Glycine. Stabilizes protein-membrane and protein-protein interactions.
Prenylation
Attachment of farnesyl or geranylgeranyl groups to C-terminal Cysteine (CaaX motif). Essential for Ras superfamily function and oncogenesis.
The Bottleneck: Why Computational Prediction?
Experimental identification (e.g., metabolic labeling, ABE, mass spec) is laborious, costly, and requires large sample inputs.
Hydrophobic nature of lipid modifications leads to poor ionization and detection difficulties in standard proteomics.
Machine learning offers a high-throughput alternative to screen the proteome for candidate substrates.
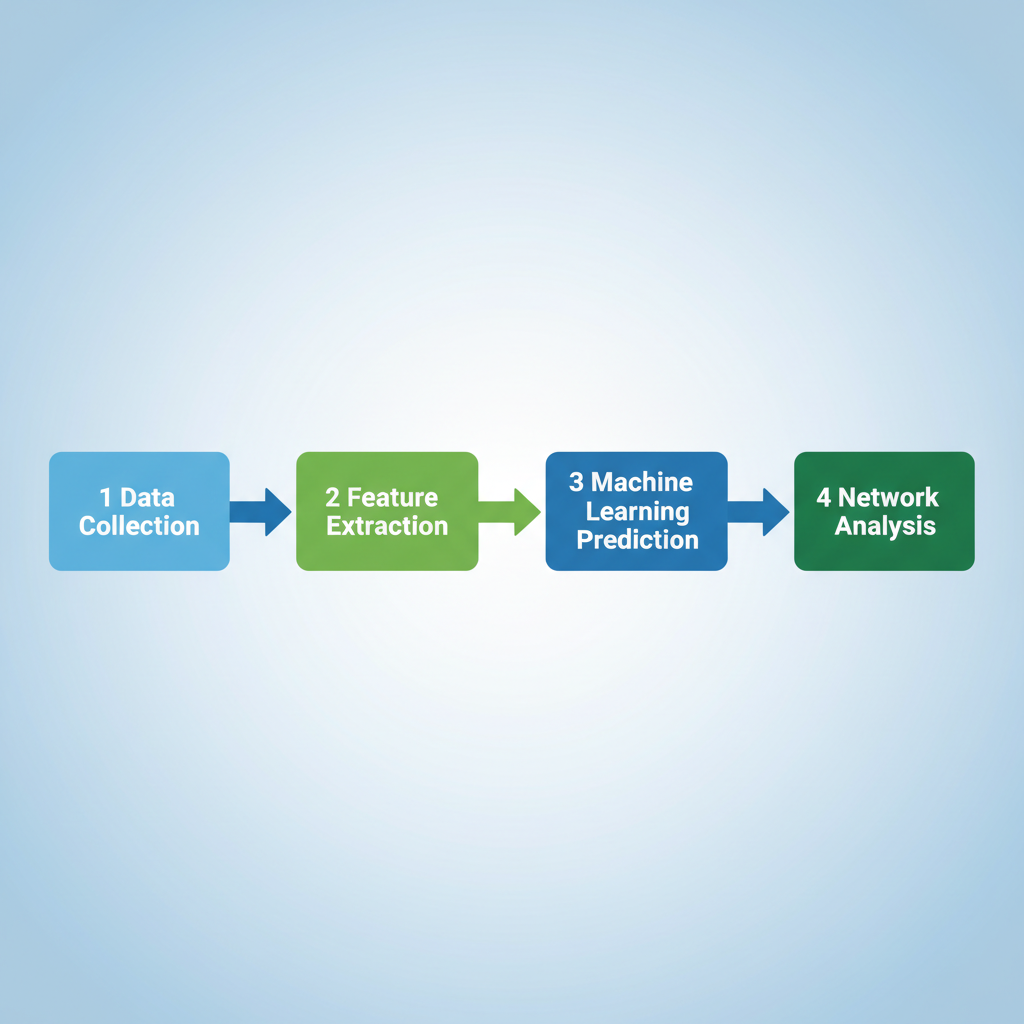
End-to-End Computational Pipeline
Data Collection: Curation of positive hits from UniProt & SwissPalm.
Feature Extraction: PseAAC, physicochemical props, structure motifs.
Model Architecture: GPS-Palm, iPrenyl, SVM/Random Forest classifiers.
Network Integration: Mapping predicted hits to PPI networks (Cytoscape).
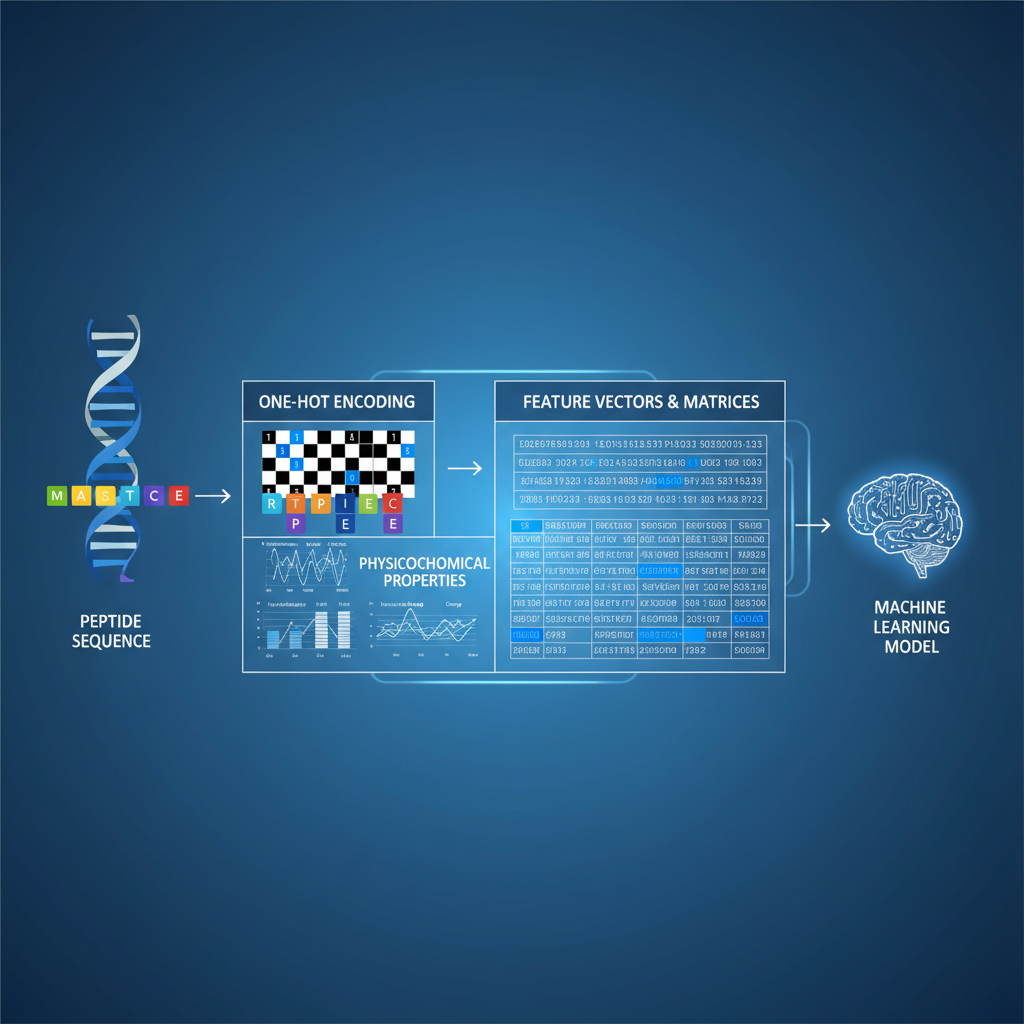
Feature Extraction Strategies
Sequence Motifs: Identification of CaaX boxes (prenylation) or Met-Gly-X-X-X-Ser/Thr (N-myristoylation).
PseAAC (Pseudo Amino Acid Composition): Incorporates sequence order information unlike standard amino acid composition.
Physicochemical Properties: Hydrophobicity, polarity, and steric hindrance around the target site.
Benchmarking S-Palmitoylation Tools
Comparison of GPS-Palm, Palmpred, and Generic ML approaches on independent test sets.
Tools for Prenylation & Myristoylation
iPrenyl-PseAAC
Utilizes Pseudo Amino Acid Composition to predict C-terminal CaaX prenylation sites. Focuses on capturing long-range sequence order effects.
PrenPs & GPS-Lipid
Employ hierarchical clustering and Group-based Prediction Systems (GPS) to reduce false positives in both N-myristoylation and Prenylation targets.
Transition to Network Biology
Predictions are just lists. Network biology contextualizes these proteins within cellular pathways.
Map predicted substrates to Protein-Protein Interaction (PPI) databases (STRING, BioGRID).
Visualize dense connectivity clusters using Cytoscape.
Functional Enrichment Analysis
Gene Ontology (GO) terms significantly enriched in the predicted lipidome.
Case Study: Disease Modules & Signaling
Constructed a specific sub-network for Ras-superfamily proteins.
Identified 'hub' proteins where prenylation inhibition disrupts oncogenic signaling cascades.
Restrained network analysis reveals cross-talk between palmitoylated receptors and downstream prenylated effectors.
Future Directions & Roadmap
Integration of Structural Data: leveraging AlphaFold models to predict solvent accessibility of Cys residues.
Deep Learning: Moving from Random Forests to Graph Neural Networks (GNNs) for better prediction accuracy.
Validation Loop: Feeding experimental validation results back into the model to refine parameters.
- bioinformatics
- protein-lipidation
- machine-learning
- network-biology
- proteomics
- cytoscape
- structural-biology