Predicting Post-Translational Modifications in Network Biology
Explore a systematic guide to PTM prediction: from data gathering and feature extraction to machine learning workflows and bioinformatics databases.
Systematic Post-Translational Modifications of Proteins
From Prediction to Network Biology
Bioinformatics & Computational Biology
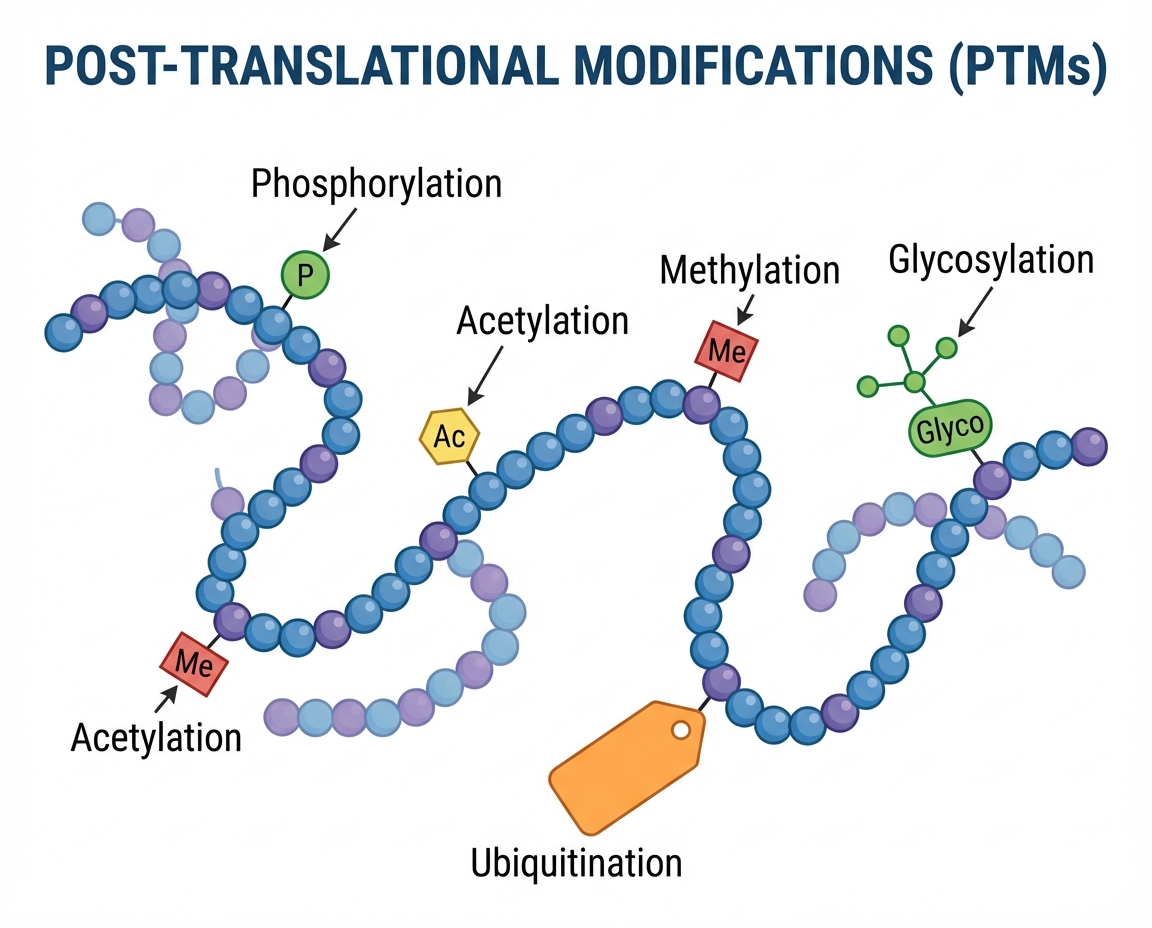
What Are Post-Translational Modifications?
Covalent processing events that change protein properties by proteolytic cleavage or adding modifying groups (acetyl, phosphoryl, glycosyl, methyl) to amino acids
Over 400 different types of PTMs identified
Can be reversible or irreversible modifications
Occur in nucleus, cytoplasm, ER, and Golgi apparatus
Affect protein structure, function, localization, and interactions
The 10 Most Studied PTM Types
Phosphorylation – Kinase-mediated, affects Ser/Thr/Tyr
Acetylation – Lysine modification, histone regulation
Ubiquitylation – Protein degradation pathway
Methylation – Epigenetic regulation on Lys/Arg
Glycosylation – N-linked and O-linked sugar chains
SUMOylation – Nuclear protein regulation
Palmitoylation – Lipid-based membrane anchoring
Myristoylation – N-terminal fatty acid addition
Prenylation – C-terminal lipid modification
Sulfation – Tyrosine modification in secreted proteins
PTM Frequency Distribution (dbPTM Database)
Top 3 PTMs comprise over 90% of all reported modification sites (~827,000 out of ~908,000)
PTM Prediction Workflow Overview
1. Data Gathering
Collect positive and negative samples from PTM databases
2. Feature Extraction
Encode sequences using biological descriptors
3. Model Training
Train classifier using machine learning algorithms
4. Performance Assessment
Evaluate using k-fold cross validation
Step 1: Data Gathering Strategies
Positive Data Selection
Sequences with experimentally verified PTM sites from databases like dbPTM or UniProt
Negative Data Selection Strategies
Strategy 1: Random proteins with target residues not undergoing PTM
Strategy 2: Proteins where no target residues have experimental PTM evidence
Strategy 3: Non-modified target residues within positive dataset proteins
Dataset Filtering
Remove redundant sequences using CD-hit (identity threshold 40-100%)
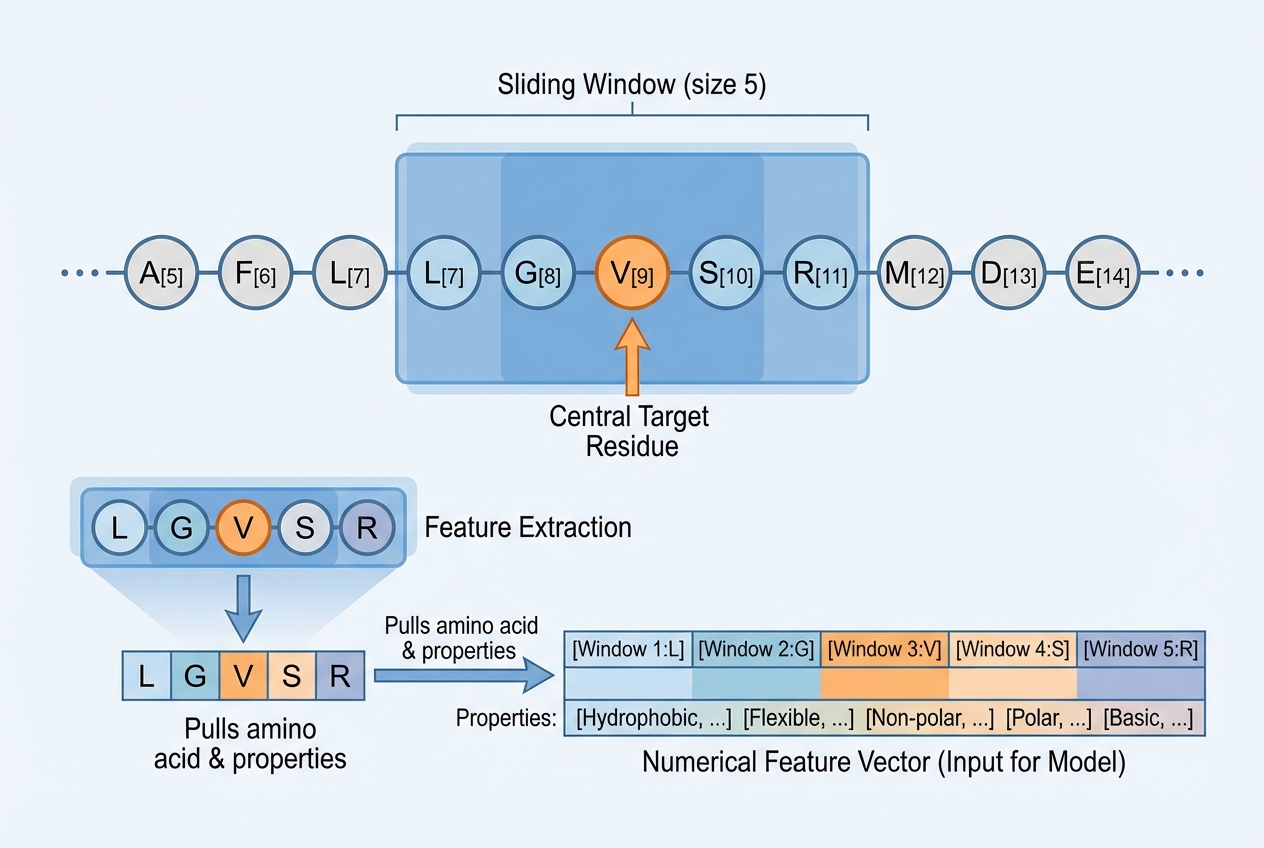
Step 2: Feature Extraction Methods
Sliding Window Approach
Partition proteins into polypeptides of length W with target residue at center. Window size typically ranges from 11 to 27 residues.
Amino Acid Composition (AAC)
Di-peptide Composition
Physicochemical Properties
Sequence Motif Similarity Scores
Position-Specific Scoring Matrices
Structural Features (secondary structure)
Major PTM Databases
dbPTM covers 130+ PTM types across 1000+ organisms; EPSD focuses on phosphorylation with 1.6M+ sites
Key PTM Prediction Tools
MusiteDeep
Deep learning for multiple PTMs (phosphorylation, glycosylation, acetylation, methylation, ubiquitination)
GPS Suite
Group-based prediction system for phosphorylation, SUMOylation, palmitoylation, methylation
NetOGlyc / NetNGlyc
Neural network-based glycosylation site prediction tools
DeepUbi / iUbiq-Lys
Specialized ubiquitination site predictors with high accuracy
PhosphoSitePlus
Comprehensive database with integrated prediction capabilities
Performance Assessment Metrics
Sensitivity (Recall)
TP / (TP + FN)
Ability to correctly identify PTM sites
Specificity
TN / (TN + FP)
Ability to correctly identify non-PTM sites
Accuracy
(TP + TN) / Total
Overall correctness of predictions
MCC
Matthews Correlation Coefficient - balanced measure for imbalanced datasets
AUC-ROC
Area Under ROC Curve - threshold-independent performance measure
PTMs in Disease and Biological Processes
Associated Diseases
Neurodegenerative diseases (Alzheimer's, Parkinson's, Huntington's)
Various cancers and metabolic syndromes
Cardiovascular and inflammatory disorders
Diabetes and immune system diseases
Key Biological Processes
Apoptosis and cell cycle control
Signal transduction and protein-protein interactions
DNA repair and transcription regulation
Chromatin organization and protein degradation
Practical Workflow: From Sequence to PTM Prediction
1. Obtain protein sequence (FASTA format from UniProt)
2. Select appropriate prediction tool based on PTM type
3. Submit sequence to web server or run local tool
4. Analyze predictions with confidence scores
5. Cross-validate with multiple tools for reliability
6. Integrate results into network/pathway analysis
Always validate computational predictions with experimental data when possible
Challenges in PTM Prediction
Data Imbalance
Negative samples often vastly outnumber positive samples, causing classifier bias
Negative Sample Selection
Absence of evidence is not evidence of absence - unlabeled sites may be undiscovered PTMs
Cross-Study Comparison
Different datasets and validation procedures make fair comparison difficult
Parameter Tuning Bias
Using same data for tuning and evaluation leads to overestimated performance
Key Takeaways
PTMs are crucial regulatory mechanisms affecting protein function, localization, and interactions
Computational prediction tools are essential due to high cost of experimental identification
Systematic workflow: Data gathering → Feature extraction → Model training → Validation
Multiple databases and tools available - choose based on PTM type and organism
Always validate predictions and be aware of methodological limitations
PTM networks provide insights into disease mechanisms and biological processes
Thank You
Questions & Discussion
dbPTM: http://dbptm.mbc.nctu.edu.tw
PhosphoSitePlus: https://www.phosphosite.org
Reference: Ramazi & Zahiri (2021) Database
- bioinformatics
- post-translational-modifications
- computational-biology
- machine-learning
- proteomics
- ptm-prediction
- network-biology