Network Biology of Protein Lipidation & Therapeutic Insights
Explore myristoylation, palmitoylation, and prenylation mechanisms. Learn how protein lipidation impacts cellular signaling and drug discovery targets.
Network Biology of Protein Lipidation
From Definitions to Systems-Level Applications
Graduate Seminar Series: Cellular Signaling & Integromics
Defining the Lipid-Modified Proteome
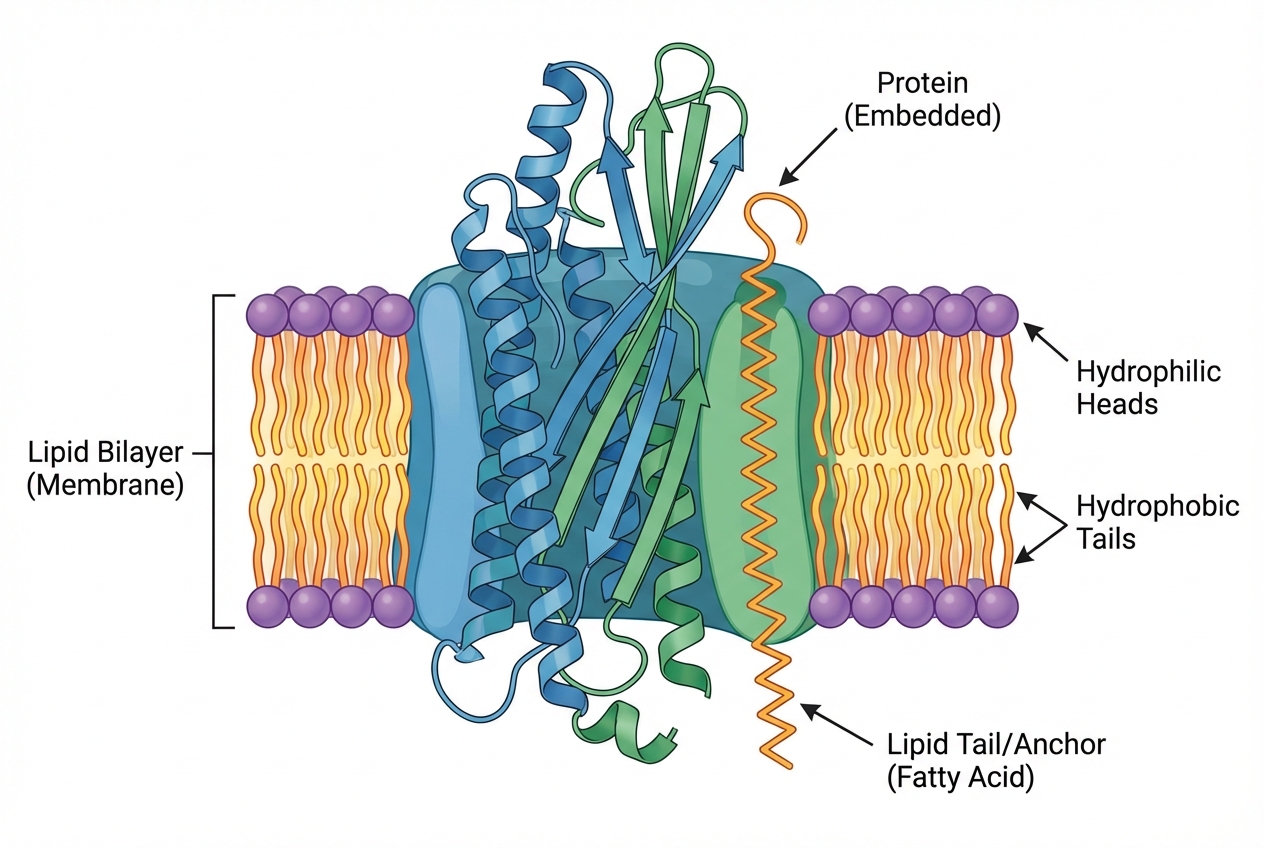
<b>Definition:</b> Covalent attachment of lipid moieties to specific amino acid residues, altering hydrophobicity and membrane affinity.
<b>Key Functions:</b> Membrane anchoring, subcellular trafficking, protein stability, and mediating Protein-Protein Interactions (PPIs).
<b>Timing:</b> Can be co-translational (e.g., N-myristoylation) or post-translational (e.g., S-palmitoylation).
Taxonomy of Protein Lipidation
<h3 style='margin:0; color:#E74C3C;'>N-Myristoylation</h3><p><b>Lipid:</b> Myristate (C14:0)<br><b>Target:</b> N-term Glycine<br><b>Nature:</b> Irreversible, Co-translational.</p>
<h3 style='margin:0; color:#27AE60;'>S-Palmitoylation</h3><p><b>Lipid:</b> Palmitate (C16:0)<br><b>Target:</b> Internal Cysteine<br><b>Nature:</b> Reversible, Post-translational.</p>
<h3 style='margin:0; color:#8E44AD;'>Prenylation</h3><p><b>Lipid:</b> Farnesyl (C15) / Geranylgeranyl (C20)<br><b>Target:</b> C-term Cysteine (CaaX box)<br><b>Nature:</b> Irreversible, Post-translational.</p>
<h3 style='margin:0; color:#F39C12;'>GPI Anchors</h3><p><b>Lipid:</b> Phosphatidylinositol glycan<br><b>Target:</b> C-terminus<br><b>Nature:</b> Complex, Luminal/Extracellular facing.</p>
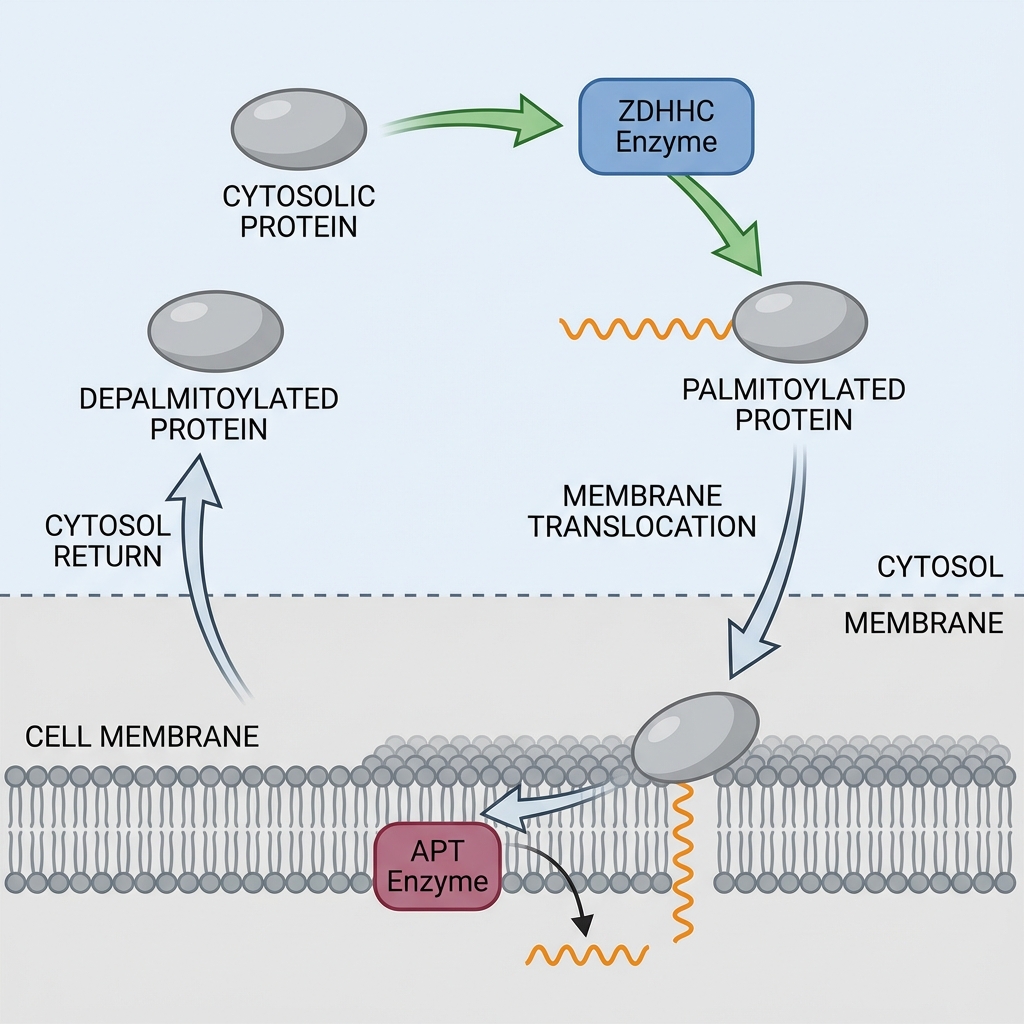
Focus: The Dynamic Palmitoylation Cycle
Unlike other lipidations, S-palmitoylation is <b>reversible</b> utilizing a thioester bond.<br><br><b>Writers:</b> ZDHHC family (PATs) - Zinc finger domain containing enzymes.<br><br><b>Erasers:</b> APTs / ABHDs (Thioesterases).<br><br>This cycle creates a rapid regulatory switch analogous to phosphorylation.
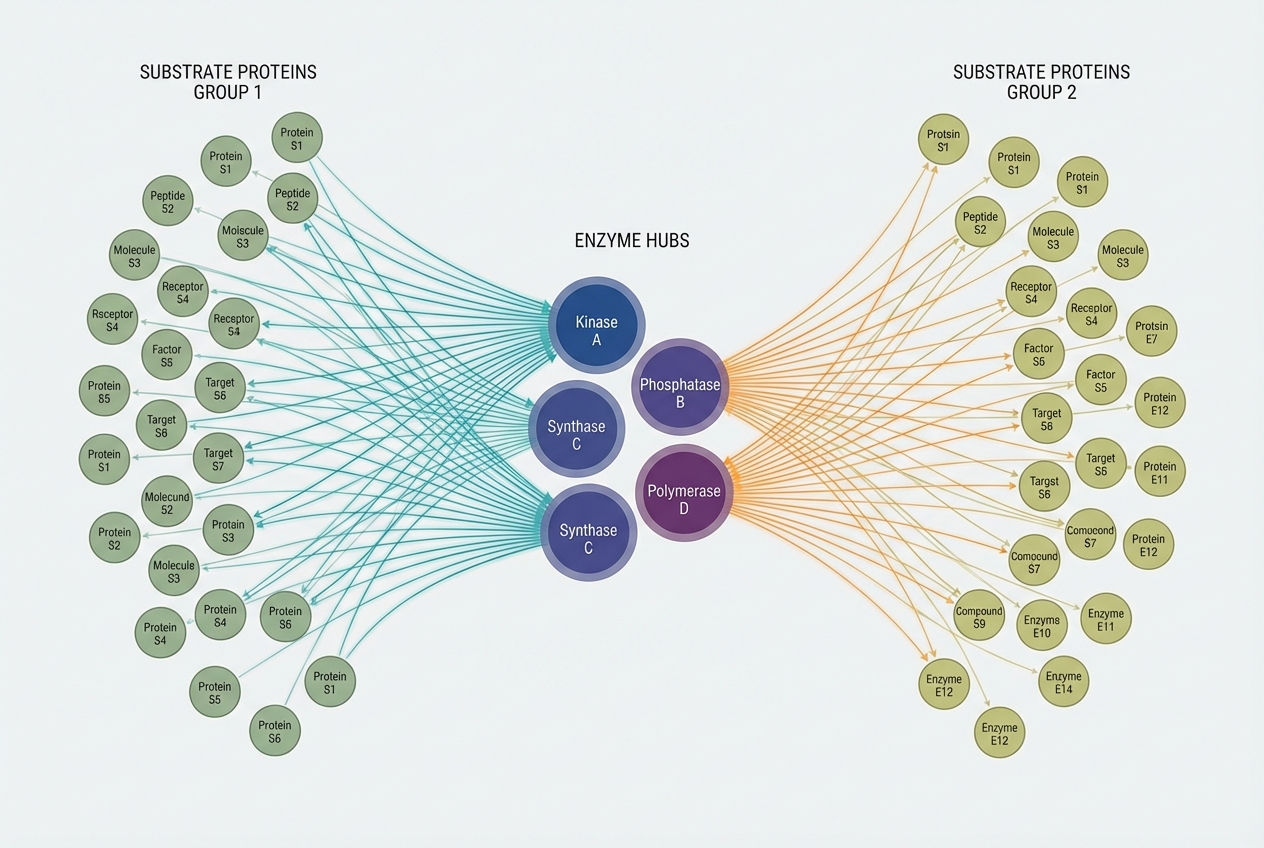
Network Biology: Enzyme-Substrate Architectures
Protein lipidation enzymes (e.g., the 23 mammalian ZDHHCs) exhibit complex substrate specificities, creating dense interaction networks.
<b>Hubs & Spokes:</b> ZDHHC enzymes act as 'hubs' modulating the localization of hundreds of 'spoke' substrates.<br><b>Redundancy:</b> Overlap in enzyme specificity provides system robustness.
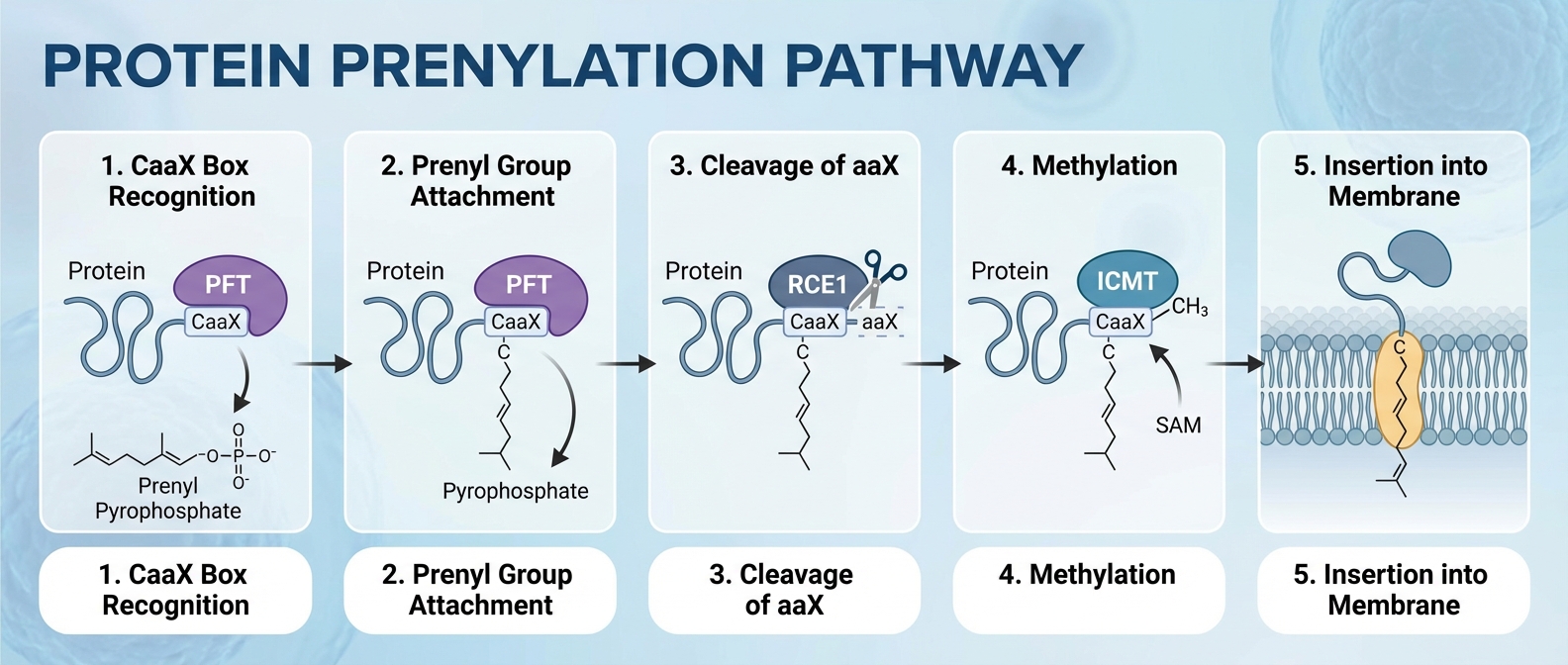
Mechanism of Prenylation: The CaaX Box
<b>Signal Motif:</b> The CaaX motif at the C-terminus.<br>C = Cysteine<br>a = Aliphatic AA<br>X = Determinant (Ser/Met -> Farnesyl; Leu -> Geranylgeranyl).
<b>Sequential Processing:</b><br>1. Cytosolic prenylation (FTase/GGTase)<br>2. ER-associated cleavage (-aaX removal)<br>3. Carboxyl-methylation (ICMT)<br>4. Result: Hydrophobic C-terminus anchors to membranes.
Membrane Compartmentalization & Trafficking
Relative distribution of lipidated proteins across cellular compartments.
<b>Kinetic Trapping:</b> Lipid modifications act as 'zip codes'.<br>• <i>Golgi:</i> Primary site of PAT activity.<br>• <i>PM:</i> Destination for signaling.<br>• <i>Acylation Cycle:</i> Allows proteins to 'hop' between internal membranes and the PM based on their lipidation state.
Case Study: The Ras Superfamily
<ul><li><b>H-Ras / N-Ras:</b> undergo both Farnesylation and Palmitoylation.</li><li><b>K-Ras:</b> undergoes Farnesylation and has a Polybasic electrostatic domain (no palmitoylation).</li><li><b>Significance:</b> 30% of human cancers are driven by Ras mutations.</li><li><b>Network Impact:</b> Lipid anchors dictate Ras isoform-specific signaling nanodomains on the PM.</li></ul>
Multi-Omics: Integrating the Lipidome
Evolution of S-Palmitoylome Detection
Modern network biology relies on high-throughput identification.<br><br><b>Integration Strategy:</b><br>1. <b>Proteomics:</b> Identify modified cysteines.<br>2. <b>Transcriptomics:</b> Correlate with ZDHHC expression profiles.<br>3. <b>Interactomics:</b> Map dynamic PPIs dependent on lipidation.
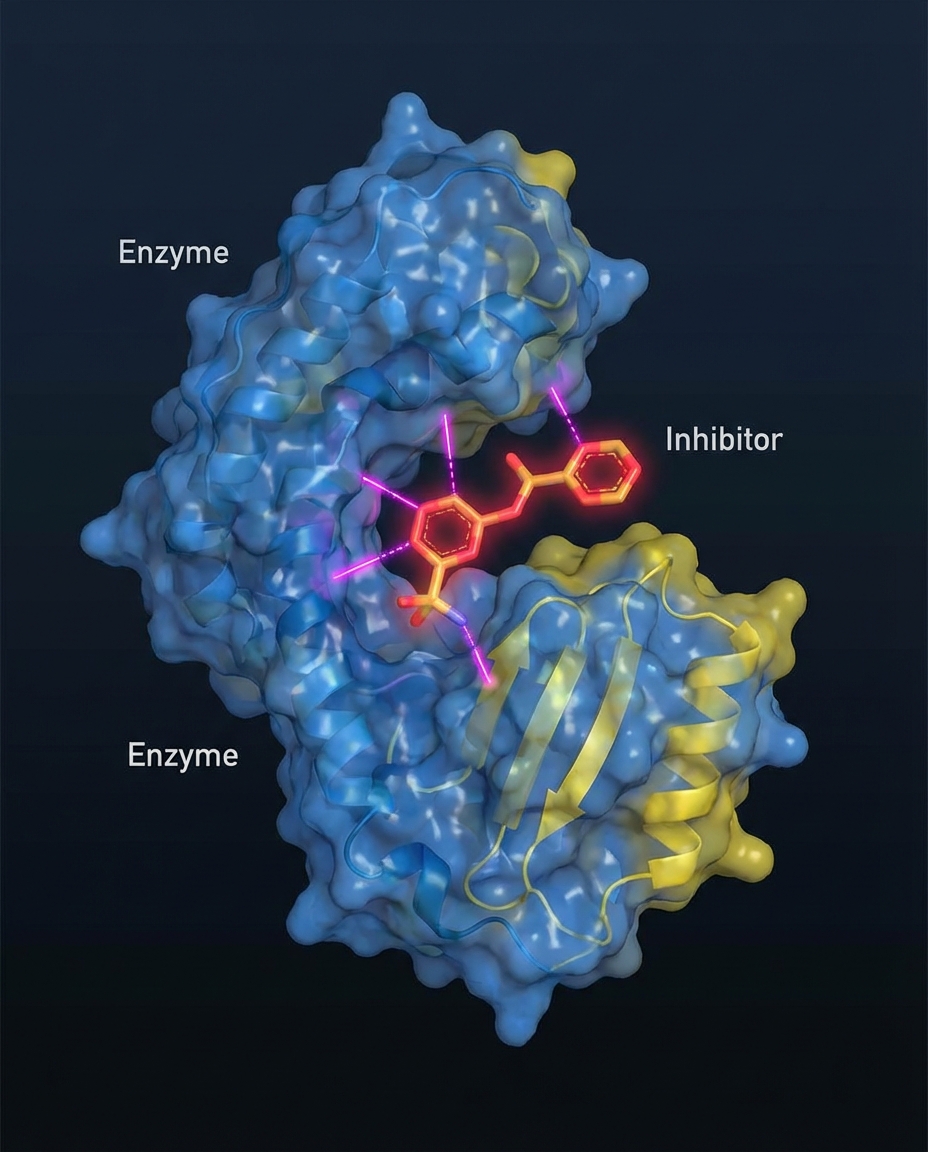
Applications: Drug Discovery Targets
<b>FTase Inhibitors (FTIs):</b><br>Initially developed for Ras-driven cancers. Revival in Progeria treatment (Lonafarnib).
<b>NMT Inhibitors:</b><br>Targeting N-myristoyltransferase in fungal infections and malaria (Plasmodium NMT).
<b>ZDHHC Specific Inhibitors:</b><br>Emerging field. Targeting specific PATs relevant to neurological disorders (e.g., ZDHHC9 in epilepsy) and cancer.
Experimental Tools: Probing the Lipidome
<b>Acyl-Biotin Exchange (ABE) / APE:</b><br>Chemical substitution of thioester-linked lipids with biotin, followed by streptavidin pull-down. <br><i>Pros:</i> No metabolic labeling needed.<br><i>Cons:</i> False positives possible.
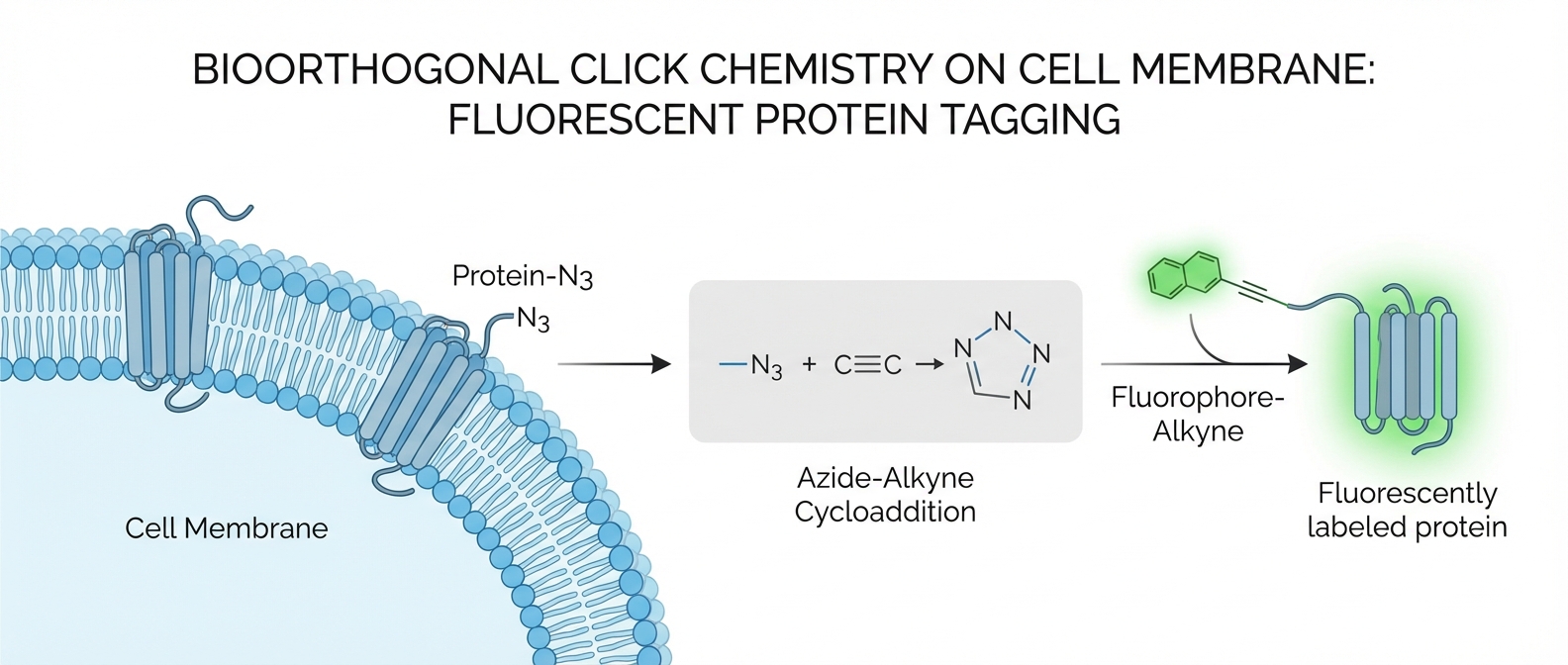
<b>Bioorthogonal 'Click' Chemistry:</b><br>Metabolic labeling with fatty acid analogs (e.g., 17-ODYA, Alk-16). Reaction with fluorescent or biotin tags.<br><i>Pros:</i> Live cell tracking, high specificity.<br><i>Cons:</i> Requires time for incorporation.
Summary & Future Directions
Protein lipidation is a critical determinant of network topology, governing subcellular localization and signaling duration.
Enzyme-substrate specificity (ZDHHCs, FTase) remains a complex 'many-to-many' network problem requiring systems biology approaches.
<b>Future Frontier:</b> Deciphering the regulating logic of the 'Palmitoylation Code' and developing isoform-specific inhibitors for clinical applications.
- protein-lipidation
- biochemistry
- cell-signaling
- systems-biology
- pharmacology
- molecular-biology
- proteomics