RNA-Mediated Safequards in CRISPR-Cas Evolution
Discover how Toxin-Antitoxin RNA pairs like the CreTA module act as addiction modules to prevent the evolutionary loss of CRISPR-Cas systems in prokaryotes.
Toxin–Antitoxin RNA Pairs Safeguard CRISPR-Cas Systems
Li et al., Science (2021) | Reviewed by [Presenter Name]
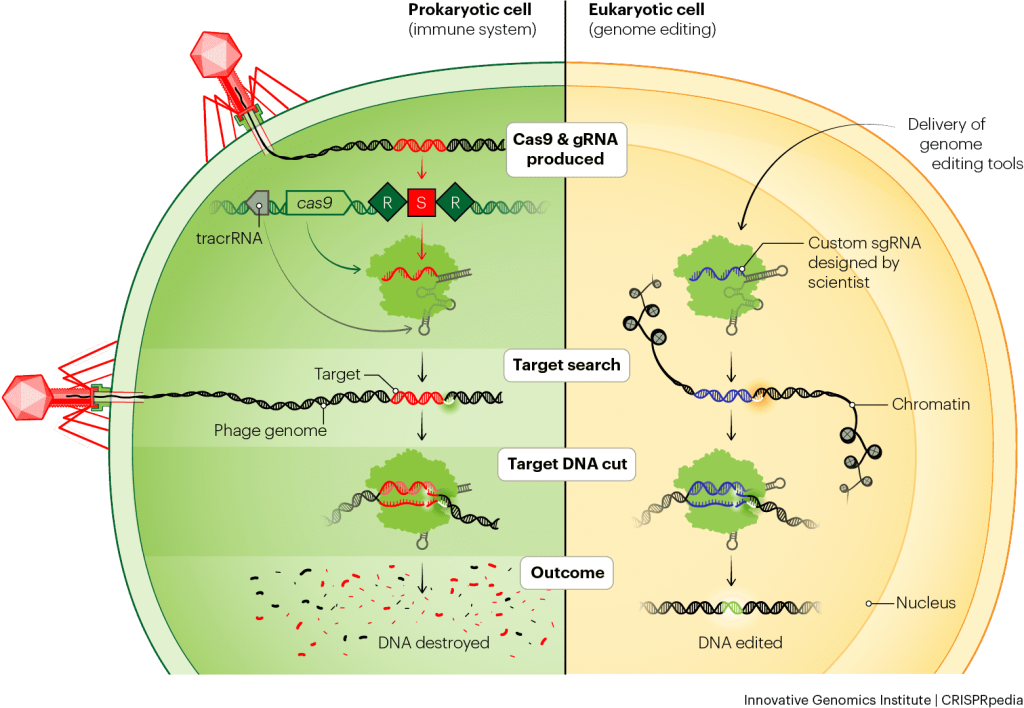
Background: RNA-Guided Adaptive Immunity
<ul><li>RNA-guided adaptive immunity mechanism</li><li>Targets invading viral DNA or RNA via complementary pairing</li><li>Provides essential antiviral defense for prokaryotes</li></ul>
The Cost of Immunity
<ul><li style='margin-bottom:30px'><strong>Autoimmunity:</strong> Risk of self-targeting host genome</li><li style='margin-bottom:30px'><strong>Genetic Conflict:</strong> Repulsion of beneficial mobile genetic elements</li><li style='margin-bottom:30px'><strong>Toxicity:</strong> Nuclease activity creates cellular stress</li><li style='margin-bottom:30px'><strong>Result:</strong> Frequent evolutionary loss of CRISPR loci</li></ul>
The Paradox: Prevalence vs. Cost
Despite high fitness costs, CRISPR-Cas systems are abundant in prokaryotes.
Hypothesis: Addiction Modules
CRISPR-Cas loci may maximize their retention by behaving as 'Selfish Genetic Elements'.
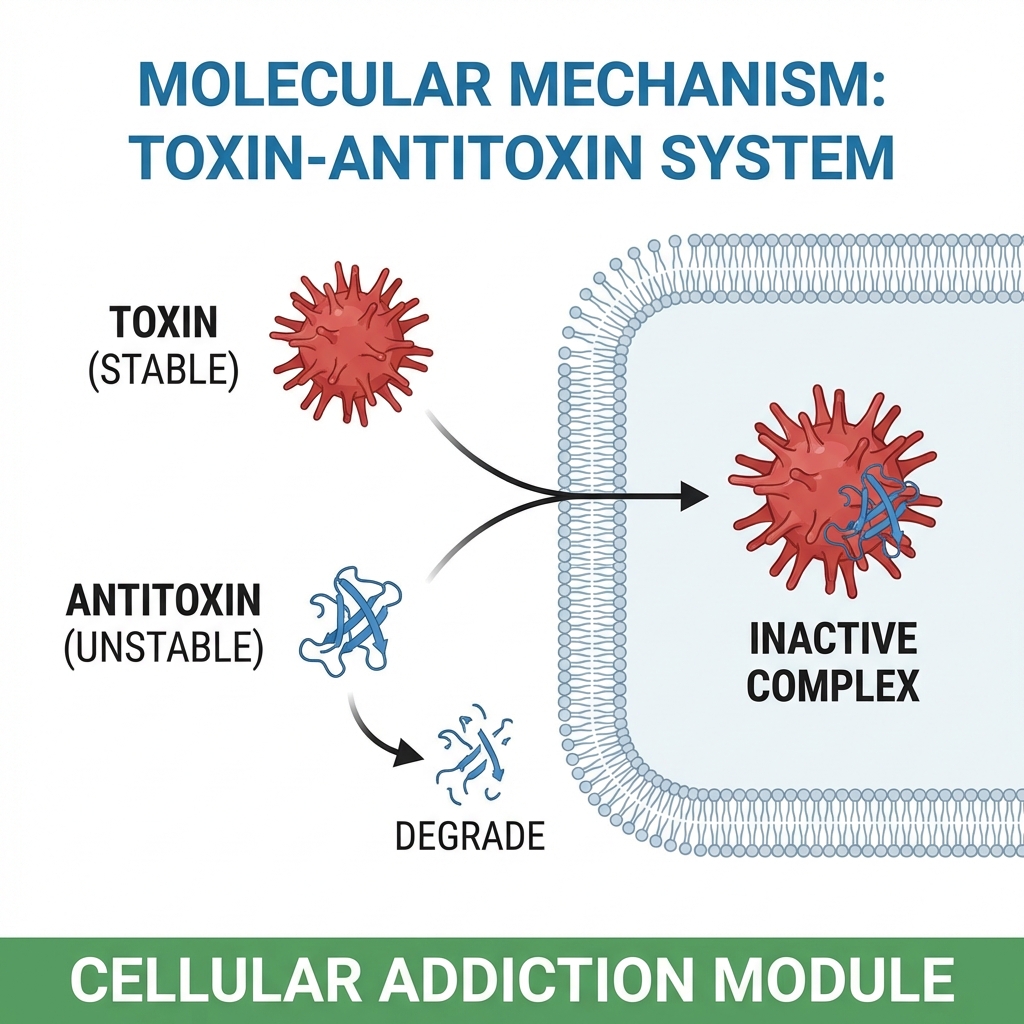
<div style='background:rgba(56, 189, 248, 0.1); padding:30px; border-left:5px solid #38bdf8; margin-bottom:20px'><strong>1. The Safeguard:</strong> Locus encodes a stable toxin and an unstable antitoxin.</div><div style='background:rgba(244, 114, 182, 0.1); padding:30px; border-left:5px solid #f472b6;'><strong>2. The Trap:</strong> If CRISPR locus is lost, antitoxin degrades, toxin persists -> Cell Death/Stasis.</div>
Discovery of the CreTA Module
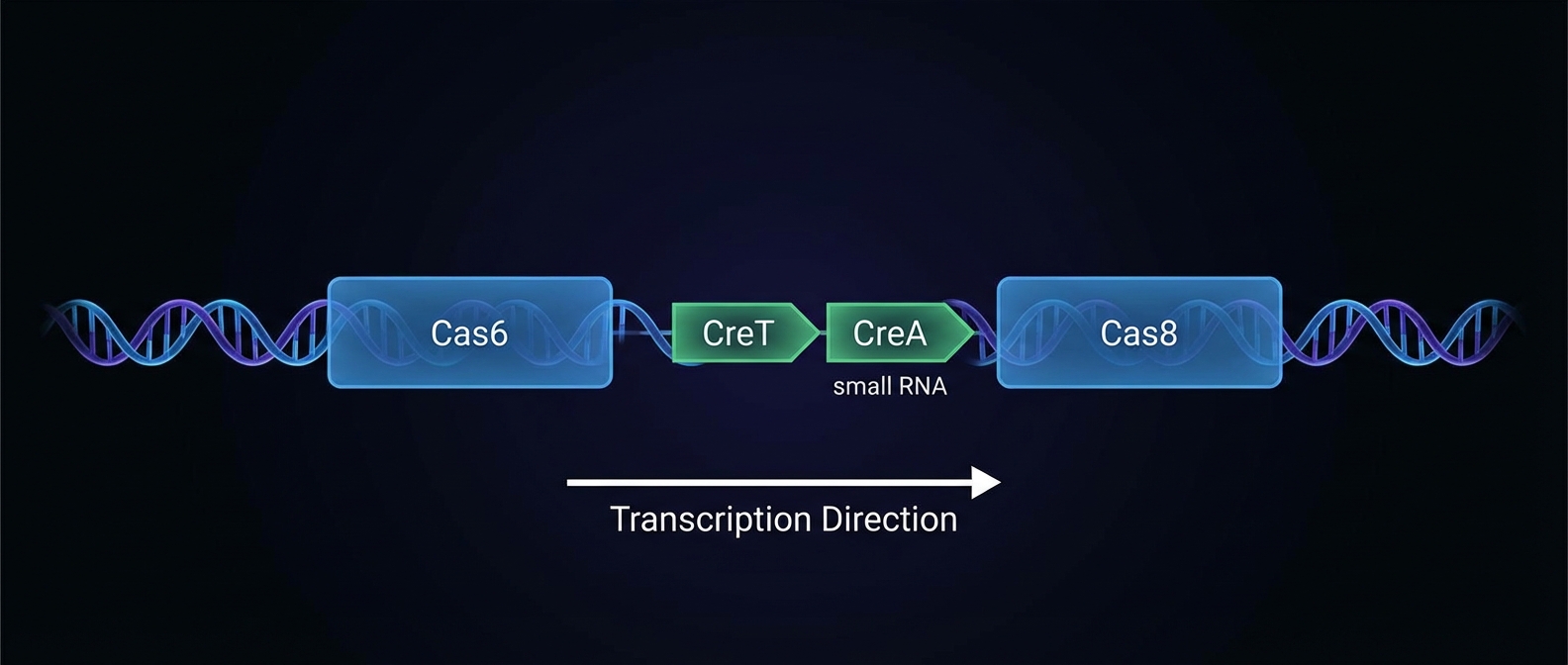
<strong>Organism:</strong> <em>Haloarcula hispanica</em><br><br><strong>System:</strong> Type I-B CRISPR-Cas<br><br><strong>Location:</strong> 311-bp intergenic region between <em>cas6</em> and <em>cas8</em> genes.<br><br><strong>Finding:</strong> A previously unidentified Toxin-Antitoxin RNA pair.
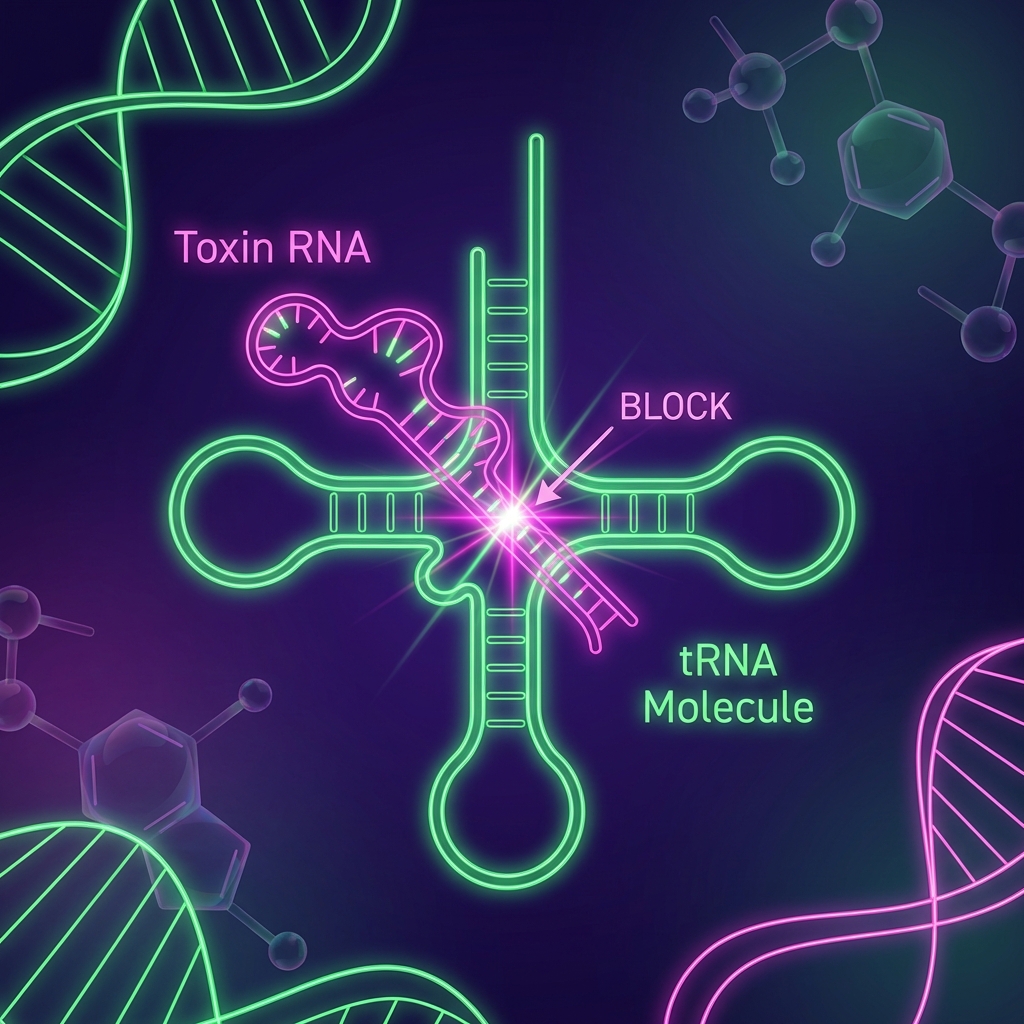
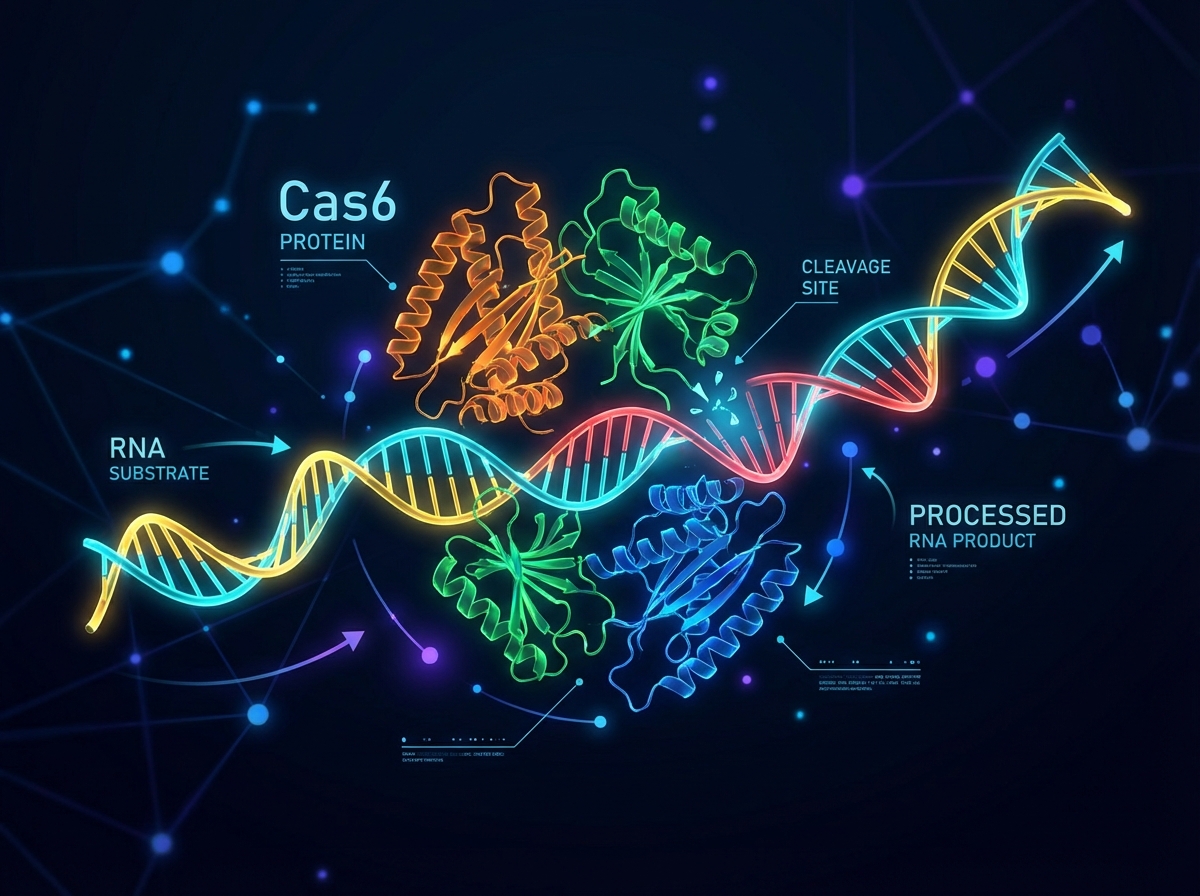
CreT: The RNA Toxin
<ul><li>Small RNA (no protein encoded)</li><li><strong>Mechanism:</strong> Sequesters rare Arginine tRNA (UCU)</li><li><strong>Effect:</strong> Impairs translation of essential genes</li><li><strong>Outcome:</strong> Bacteriostatic (arrests growth) rather than bactericidal</li></ul>
CreA: The crRNA-like Antitoxin
<ul><li>Resembles a canonical CRISPR RNA (crRNA)</li><li>Processed specifically by <strong>Cas6</strong> endonuclease</li><li>Lacks the standard 3' handle</li><li><strong>Dependence:</strong> Functional only when bound to the Cascade effector complex</li></ul>
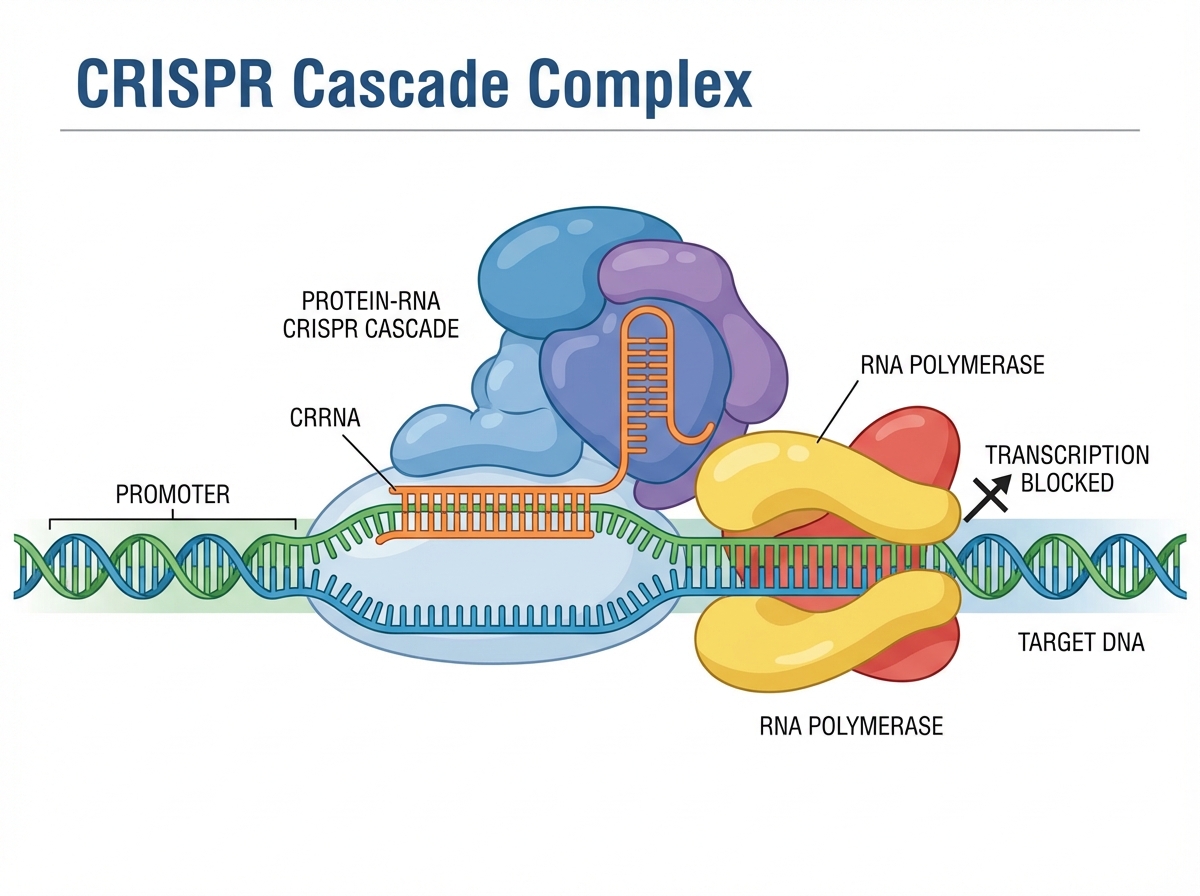
Mechanism: Repression, Not Cleavage
<ul><li>CreA guides Cascade to the <em>creT</em> promoter</li><li><strong>Partial Complementarity:</strong> PAM + Seed region match only</li><li><strong>Result:</strong> Cascade sits on DNA, physically blocking transcription</li><li><strong>Key Distinction:</strong> Functional gene silencing without degrading the DNA</li></ul>
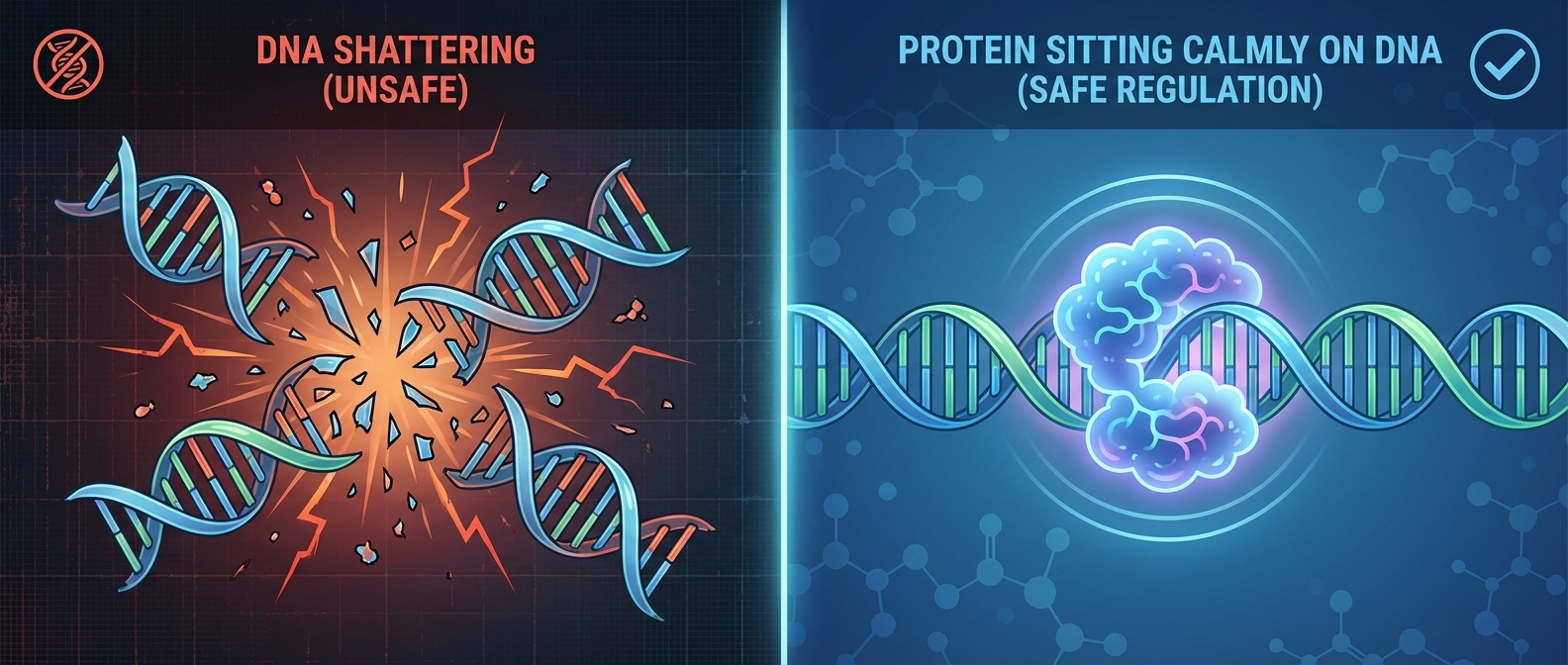
Safety: Avoiding Autoimmunity
<div style='display:flex; gap:40px; margin-top:40px;'><div style='flex:1; background:#0f172a; padding:30px; border-radius:15px; border-top:5px solid #ef4444;'><h3>Standard CRISPR</h3><p>Full complementarity guides <strong>Cas3</strong> recruitment causing massive DNA degradation.</p></div><div style='flex:1; background:#0f172a; padding:30px; border-radius:15px; border-top:5px solid #22c55e;'><h3>CreTA System</h3><p>Partial matching prevents Cas3 recruitment. DNA remains intact (Safe Regulation).</p></div></div>
- crispr-cas
- molecular-biology
- genetics
- rna-toxin
- biotechnology
- evolutionary-biology
- microbiology